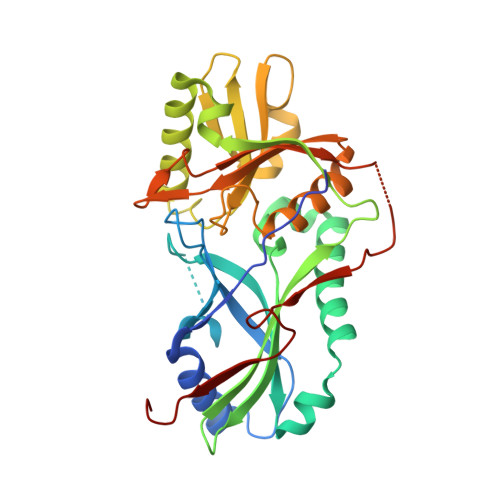
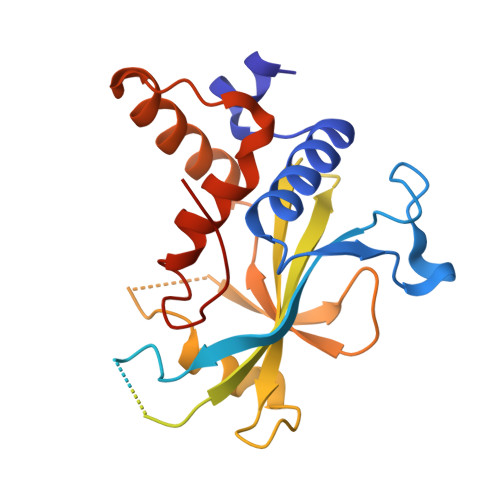
Noncanonical recognition and UBL loading of distinct E2s by autophagy-essential Atg7.
Yamaguchi, M., Matoba, K., Sawada, R., Fujioka, Y., Nakatogawa, H., Yamamoto, H., Kobashigawa, Y., Hoshida, H., Akada, R., Ohsumi, Y., Noda, N.N., Inagaki, F.(2012) Nat Struct Mol Biol 19: 1250-1256
- PubMed: 23142983 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2451
- Primary Citation Related Structures:
3VX6, 3VX7, 3VX8 - PubMed Abstract:
Autophagy requires ubiquitin-like Atg8 and Atg12 conjugation systems, where Atg7 has a critical role as the sole E1 enzyme. Although Atg7 recognizes two distinct E2s, Atg3 and Atg10, it is not understood how Atg7 correctly loads these E2s with their cognate ubiquitin-like proteins, Atg8 and Atg12. Here, we report the crystal structures of the N-terminal domain of Atg7 bound to Atg10 or Atg3 of thermotolerant yeast and plant homologs. The observed Atg7-Atg10 and Atg7-Atg3 interactions, which resemble each other but are quite distinct from the canonical E1-E2 interaction, makes Atg7 suitable for transferring Atg12 to Atg10 and Atg8 to Atg3 by a trans mechanism. Notably, in vitro experiments showed that Atg7 loads Atg3 and Atg10 with Atg8 and Atg12 in a nonspecific manner, which suggests that cognate conjugate formation in vivo is not an intrinsic quality of Atg7.
- Department of Structural Biology, Faculty of Advanced Life Science, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: