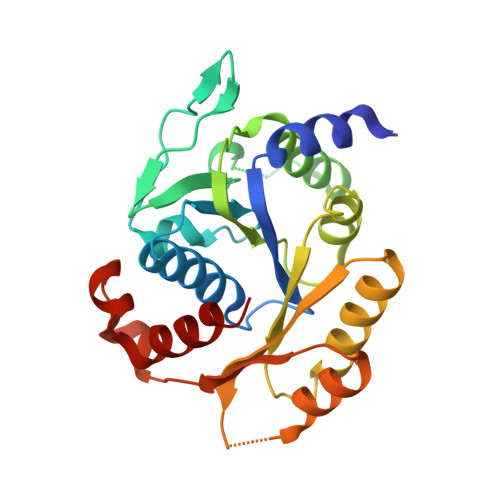
Identification and Structure of a Novel Archaeal HypB for [NiFe] Hydrogenase Maturation
Sasaki, D., Watanabe, S., Matsumi, R., Shoji, T., Yasukochi, A., Tagashira, K., Fukuda, W., Kanai, T., Atomi, H., Imanaka, T., Miki, K.(2013) J Mol Biology 425: 1627-1640
- PubMed: 23399544 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.02.004
- Primary Citation Related Structures:
3VX3 - PubMed Abstract:
HypB (metal-binding GTPase) and HypA (nickel metallochaperone) are required for nickel insertion into [NiFe] hydrogenase. However, the HypB homolog proteins are not found in some archaeal species including Thermococcales. In this article, we identify a novel archaeal Mrp/MinD family ATPase-type HypB from Thermococcus kodakarensis (Tk-mmHypB) and determine its crystal structure. The mmhypB gene is conserved among species lacking the hypB gene and is located adjacent to the hypA gene on their genome. Deletion of the mmhypB gene leads to a significant reduction in hydrogen-dependent growth of T. kodakarensis, which is restored by nickel supplementation. The monomer structure of Tk-mmHypB is similar to those of the Mrp/MinD family ATPases. The ADP molecules are tightly bound to the protein. Isothermal titration calorimetry shows that Tk-mmHypB binds ATP with a K(d) value of 84 nM. ADP binds more tightly than does ATP, with a K(d) value of 15 nM. The closed Tk-mmHypB dimer in the crystallographic asymmetric unit is consistent with the ATP-hydrolysis-deficient dimer of the Mrp/MinD family Soj/MinD proteins. Structural comparisons with these proteins suggest the ATP-binding dependent conformational change and rearrangement of the Tk-mmHypB dimer. These observations imply that the nickel insertion process during the [NiFe] hydrogenase maturation is performed by HypA, mmHypB, and a nucleotide exchange factor in these archaea.
- Department of Chemistry, Graduate School of Science, Kyoto University, Kyoto 606-8502, Japan.
Organizational Affiliation: