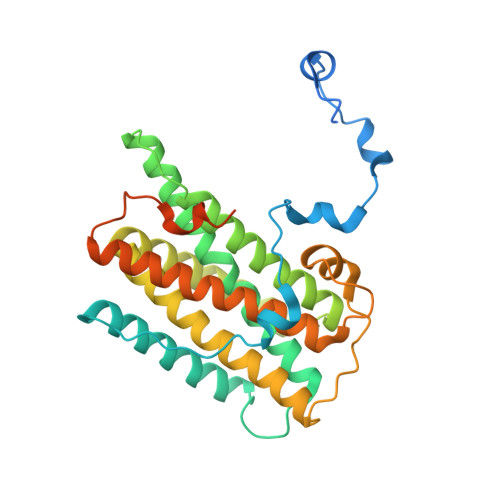
Structure of the trypanosome cyanide-insensitive alternative oxidase
Shiba, T., Kido, Y., Sakamoto, K., Inaoka, D.K., Tsuge, C., Tatsumi, R., Takahashi, G., Balogun, E.O., Nara, T., Aoki, T., Honma, T., Tanaka, A., Inoue, M., Matsuoka, S., Saimoto, H., Moore, A.L., Harada, S., Kita, K.(2013) Proc Natl Acad Sci U S A 110: 4580-4585
- PubMed: 23487766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1218386110
- Primary Citation Related Structures:
3VV9, 3VVA, 3W54 - PubMed Abstract:
In addition to haem copper oxidases, all higher plants, some algae, yeasts, molds, metazoans, and pathogenic microorganisms such as Trypanosoma brucei contain an additional terminal oxidase, the cyanide-insensitive alternative oxidase (AOX). AOX is a diiron carboxylate protein that catalyzes the four-electron reduction of dioxygen to water by ubiquinol. In T. brucei, a parasite that causes human African sleeping sickness, AOX plays a critical role in the survival of the parasite in its bloodstream form. Because AOX is absent from mammals, this protein represents a unique and promising therapeutic target. Despite its bioenergetic and medical importance, however, structural features of any AOX are yet to be elucidated. Here we report crystal structures of the trypanosomal alternative oxidase in the absence and presence of ascofuranone derivatives. All structures reveal that the oxidase is a homodimer with the nonhaem diiron carboxylate active site buried within a four-helix bundle. Unusually, the active site is ligated solely by four glutamate residues in its oxidized inhibitor-free state; however, inhibitor binding induces the ligation of a histidine residue. A highly conserved Tyr220 is within 4 Å of the active site and is critical for catalytic activity. All structures also reveal that there are two hydrophobic cavities per monomer. Both inhibitors bind to one cavity within 4 Å and 5 Å of the active site and Tyr220, respectively. A second cavity interacts with the inhibitor-binding cavity at the diiron center. We suggest that both cavities bind ubiquinol and along with Tyr220 are required for the catalytic cycle for O2 reduction.
- Department of Biomedical Chemistry, Graduate School of Medicine, The University of Tokyo, Tokyo 113-0033, Japan.
Organizational Affiliation: