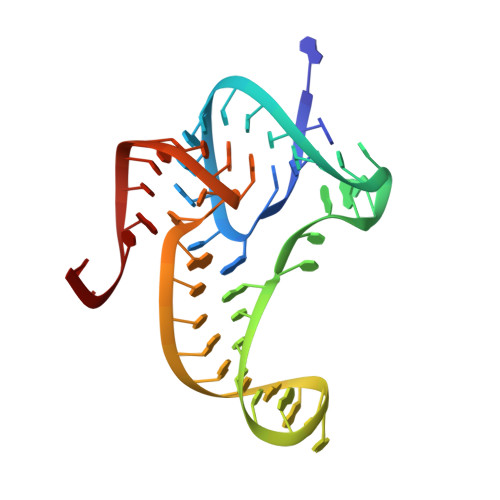
Fluoride ion encapsulation by Mg2+ ions and phosphates in a fluoride riboswitch.
Ren, A., Rajashankar, K.R., Patel, D.J.(2012) Nature 486: 85-89
- PubMed: 22678284 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11152
- Primary Citation Related Structures:
3VRS, 4EN5, 4ENA, 4ENB, 4ENC - PubMed Abstract:
Significant advances in our understanding of RNA architecture, folding and recognition have emerged from structure-function studies on riboswitches, non-coding RNAs whose sensing domains bind small ligands and whose adjacent expression platforms contain RNA elements involved in the control of gene regulation. We now report on the ligand-bound structure of the Thermotoga petrophila fluoride riboswitch, which adopts a higher-order RNA architecture stabilized by pseudoknot and long-range reversed Watson-Crick and Hoogsteen A•U pair formation. The bound fluoride ion is encapsulated within the junctional architecture, anchored in place through direct coordination to three Mg(2+) ions, which in turn are octahedrally coordinated to water molecules and five inwardly pointing backbone phosphates. Our structure of the fluoride riboswitch in the bound state shows how RNA can form a binding pocket selective for fluoride, while discriminating against larger halide ions. The T. petrophila fluoride riboswitch probably functions in gene regulation through a transcription termination mechanism.
- Structural Biology Program, Memorial Sloan-Kettering Center, New York, New York 10065, USA.
Organizational Affiliation: