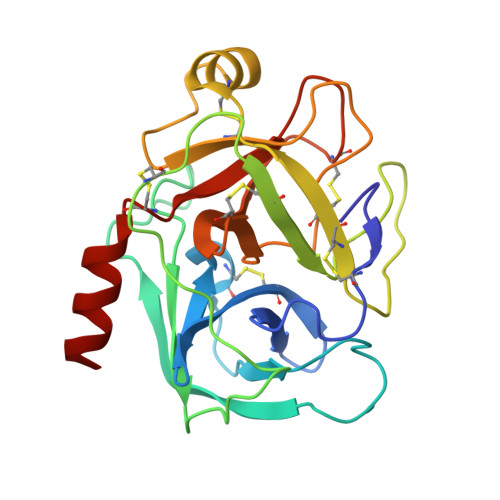
Crystal structure of 6-guanidinohexanoyl trypsin near the optimum pH reveals the acyl-enzyme intermediate to be deacylated
Masuda, Y., Nitanai, Y., Mizutani, R., Noguchi, S.(2012) Proteins
- PubMed: 23161653 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24206
- Primary Citation Related Structures:
3VPK - PubMed Abstract:
The force driving the conversion from the acyl intermediate to the tetrahedral intermediate in the deacylation reaction of serine proteases remains unclear. The crystal structure of 6-guanidinohexanoyl trypsin was determined at pH 7.0, near the optimum reaction pH, at 1.94 Å resolution. In this structure, three water molecules are observed around the catalytic site. One acts as a nucleophile to attack the acyl carbonyl carbon while the other two waters fix the position of the catalytic water through a hydrogen bond. When the acyl carbonyl oxygen oscillates thermally, the water assumes an appropriate angle to catalyze the deacylation.
- Nuclear, Biological and Chemical Detection Technology Section, Human Oriented Systems Division, Advanced Defense Technology Center, Technical Research and Development Institute, Ministry of Defense, Meguro, Tokyo 153-8630, Japan. mas@cs.trdi.mod.go.jp
Organizational Affiliation: