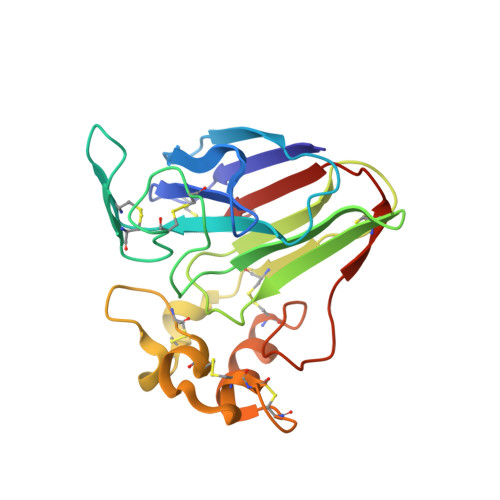
Atomic structure of the sweet-tasting protein thaumatin I at pH 8.0 reveals the large disulfide-rich region in domain II to be sensitive to a pH change
Masuda, T., Ohta, K., Mikami, B., Kitabatake, N., Tani, F.(2012) Biochem Biophys Res Commun 419: 72-76
- PubMed: 22326916 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2012.01.129
- Primary Citation Related Structures:
3VHF, 3VJQ - PubMed Abstract:
Thaumatin, an intensely sweet-tasting plant protein, elicits a sweet taste at 50 nM. Although the sweetness remains when thaumatin is heated at 80 °C for 4h under acid conditions, it rapidly declines when heating at a pH above 6.5. To clarify the structural difference at high pH, the atomic structure of a recombinant thaumatin I at pH 8.0 was determined at a resolution of 1.0Å. Comparison to the crystal structure of thaumatin at pH 7.3 and 7.0 revealed the root-mean square deviation value of a Cα atom to be substantially greater in the large disulfide-rich region of domain II, especially residues 154-164, suggesting that a loop region in domain II to be affected by solvent conditions. Furthermore, B-factors of Lys137, Lys163, and Lys187 were significantly affected by pH change, suggesting that a striking increase in the mobility of these lysine residues, which could facilitate a reaction with a free sulfhydryl residue produced via the β-elimination of disulfide bonds by heating at a pH above 7.0. The increase in mobility of lysine residues as well as a loop region in domain II might play an important role in the heat-induced aggregation of thaumatin above pH 7.0.
- Division of Food Science and Biotechnology, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan. t2masuda@kais.kyoto-u.ac.jp
Organizational Affiliation: