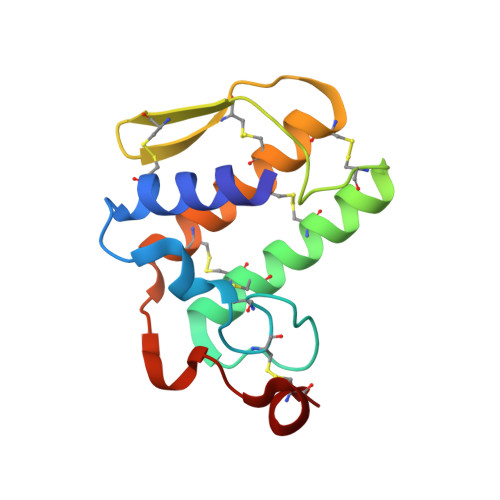
Mechanistic studies on the anticoagulant activity of a phospholipase A2 from the venom of the Australian King Brown Snake (Pseudechis australis)
Trabi, M., Millers, E.-K., Richards, R., Snelling, H., Keegan, R., Lavin, M.F., de Jersey, J., Guddat, L.W., Masci, P.To be published.