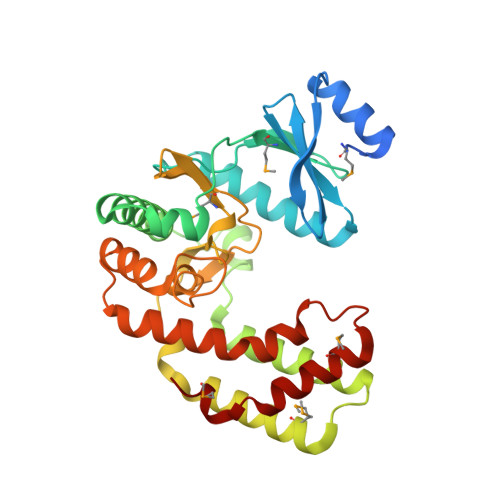
Crystal structure of aminoglycoside phosphotransferase APH(2'')-Ib, apo form
Stogios, P.J., Minasov, G., Singer, A.U., Tan, K., Nocek, B., Evdokimova, E., Egorova, E., Di Leo, R., Savchenko, A., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.