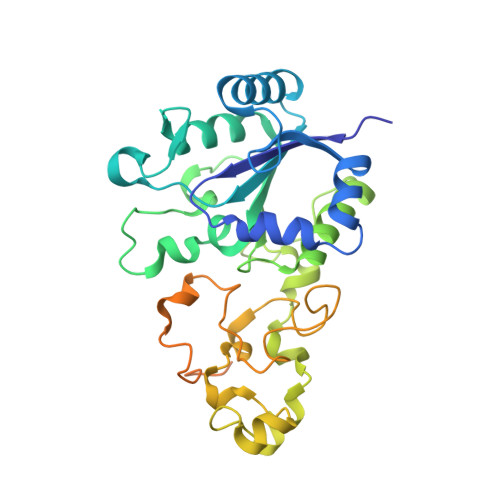
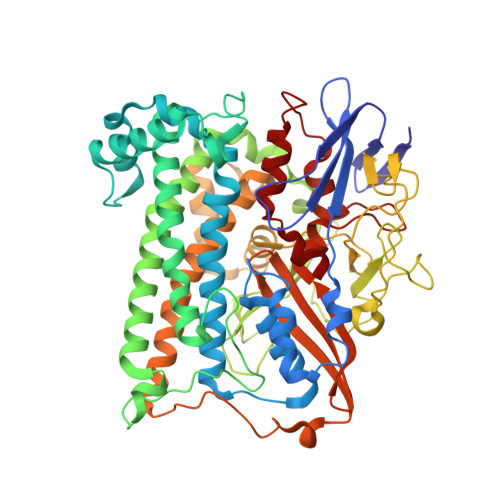
X-ray crystallographic and computational studies of the O2-tolerant [NiFe]-hydrogenase 1 from Escherichia coli.
Volbeda, A., Amara, P., Darnault, C., Mouesca, J.M., Parkin, A., Roessler, M.M., Armstrong, F.A., Fontecilla-Camps, J.C.(2012) Proc Natl Acad Sci U S A 109: 5305-5310
- PubMed: 22431599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1119806109
- Primary Citation Related Structures:
3UQY, 3USC, 3USE - PubMed Abstract:
The crystal structure of the membrane-bound O(2)-tolerant [NiFe]-hydrogenase 1 from Escherichia coli (EcHyd-1) has been solved in three different states: as-isolated, H(2)-reduced, and chemically oxidized. As very recently reported for similar enzymes from Ralstonia eutropha and Hydrogenovibrio marinus, two supernumerary Cys residues coordinate the proximal [FeS] cluster in EcHyd-1, which lacks one of the inorganic sulfide ligands. We find that the as-isolated, aerobically purified species contains a mixture of at least two conformations for one of the cluster iron ions and Glu76. In one of them, Glu76 and the iron occupy positions that are similar to those found in O(2)-sensitive [NiFe]-hydrogenases. In the other conformation, this iron binds, besides three sulfur ligands, the amide N from Cys20 and one Oε of Glu76. Our calculations show that oxidation of this unique iron generates the high-potential form of the proximal cluster. The structural rearrangement caused by oxidation is confirmed by our H(2)-reduced and oxidized EcHyd-1 structures. Thus, thanks to the peculiar coordination of the unique iron, the proximal cluster can contribute two successive electrons to secure complete reduction of O(2) to H(2)O at the active site. The two observed conformations of Glu76 are consistent with this residue playing the role of a base to deprotonate the amide moiety of Cys20 upon iron binding and transfer the resulting proton away, thus allowing the second oxidation to be electroneutral. The comparison of our structures also shows the existence of a dynamic chain of water molecules, resulting from O(2) reduction, located near the active site.
- Metalloproteins Unit, Institut de Biologie Structurale J.-P. Ebel, Commissariat à l'Energie Atomique-Centre National de la Recherche Scientifique-l'Université Joseph Fourier, 41 Rue Jules Horowitz, 38027 Grenoble, France.
Organizational Affiliation: