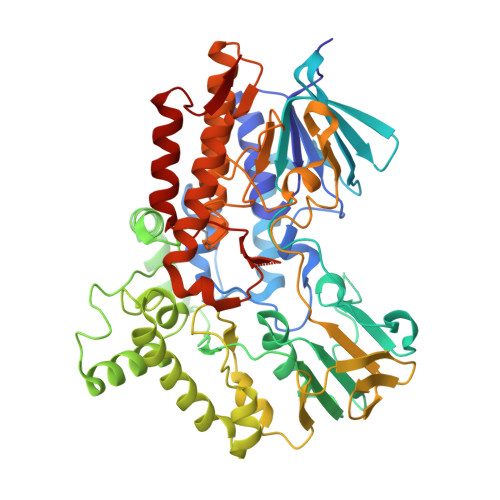
Cloning, Baeyer-Villiger biooxidations, and structures of the camphor pathway 2-oxo-{Delta}(3)-4,5,5-trimethylcyclopentenylacetyl-coenzyme A monooxygenase of Pseudomonas putida ATCC 17453.
Leisch, H., Shi, R., Grosse, S., Morley, K., Bergeron, H., Cygler, M., Iwaki, H., Hasegawa, Y., Lau, P.C.(2012) Appl Environ Microbiol 78: 2200-2212
- PubMed: 22267661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AEM.07694-11
- Primary Citation Related Structures:
3UOV, 3UOX, 3UOY, 3UOZ, 3UP4, 3UP5 - PubMed Abstract:
A dimeric Baeyer-Villiger monooxygenase (BVMO) catalyzing the lactonization of 2-oxo-Δ(3)-4,5,5-trimethylcyclopentenylacetyl-coenzyme A (CoA), a key intermediate in the metabolism of camphor by Pseudomonas putida ATCC 17453, had been initially characterized in 1983 by Ougham and coworkers (H. J. Ougham, D. G. Taylor, and P. W. Trudgill, J. Bacteriol. 153:140-152, 1983). Here we cloned and overexpressed the 2-oxo-Δ(3)-4,5,5-trimethylcyclopentenylacetyl-CoA monooxygenase (OTEMO) in Escherichia coli and determined its three-dimensional structure with bound flavin adenine dinucleotide (FAD) at a 1.95-Å resolution as well as with bound FAD and NADP(+) at a 2.0-Å resolution. OTEMO represents the first homodimeric type 1 BVMO structure bound to FAD/NADP(+). A comparison of several crystal forms of OTEMO bound to FAD and NADP(+) revealed a conformational plasticity of several loop regions, some of which have been implicated in contributing to the substrate specificity profile of structurally related BVMOs. Substrate specificity studies confirmed that the 2-oxo-Δ(3)-4,5,5-trimethylcyclopentenylacetic acid coenzyme A ester is preferred over the free acid. However, the catalytic efficiency (k(cat)/K(m)) favors 2-n-hexyl cyclopentanone (4.3 × 10(5) M(-1) s(-1)) as a substrate, although its affinity (K(m) = 32 μM) was lower than that of the CoA-activated substrate (K(m) = 18 μM). In whole-cell biotransformation experiments, OTEMO showed a unique enantiocomplementarity to the action of the prototypical cyclohexanone monooxygenase (CHMO) and appeared to be particularly useful for the oxidation of 4-substituted cyclohexanones. Overall, this work extends our understanding of the molecular structure and mechanistic complexity of the type 1 family of BVMOs and expands the catalytic repertoire of one of its original members.
- Biotechnology Research Institute, National Research Council Canada, Montreal, Quebec, Canada.
Organizational Affiliation: