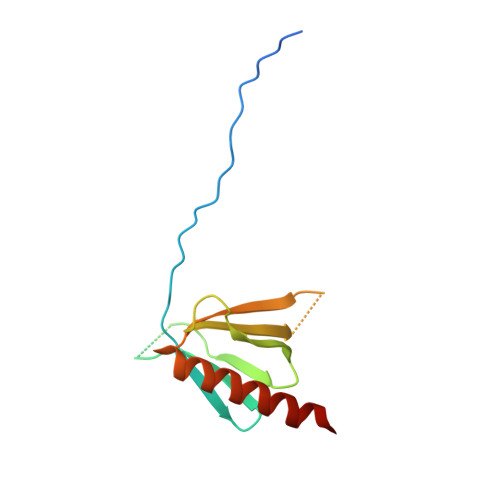
Structures of the pleckstrin homology domain of Saccharomyces cerevisiae Avo1 and its human orthologue Sin1, an essential subunit of TOR complex 2
Pan, D., Matsuura, Y.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 386-392
- PubMed: 22505404 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112007178
- Primary Citation Related Structures:
3ULB, 3ULC, 3VOQ - PubMed Abstract:
In eukaryotes, multiprotein complexes termed TOR complex 1 (TORC1) and TOR complex 2 (TORC2) function as major regulators of cell growth, metabolism and ageing. The C-terminal domain of the Saccharomyces cerevisiae TORC2 component Avo1 is required for plasma-membrane localization of TORC2 and is essential for yeast viability. X-ray crystal structures of the C-terminal domain of Avo1 and of its human orthologue Sin1 have been determined. The structures show that the C-termini of Avo1 and Sin1 both have the pleckstrin homology (PH) domain fold. Comparison with known PH-domain structures suggests a putative binding site for phosphoinositides.
- Division of Biological Science, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan.
Organizational Affiliation: