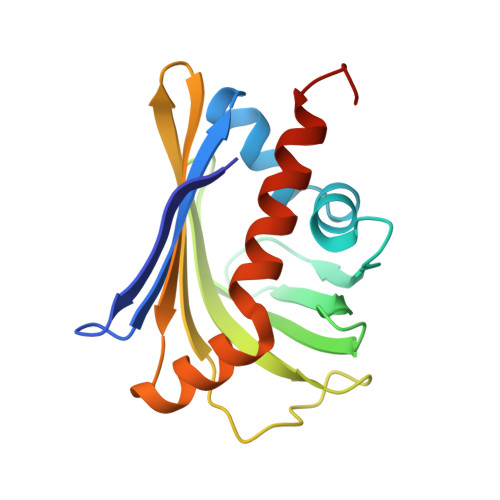
Crystal structure of the toxin Msmeg_6760, the structural homolog of Mycobacterium tuberculosis Rv2035, a novel type II toxin involved in the hypoxic response.
Bajaj, R.A., Arbing, M.A., Shin, A., Cascio, D., Miallau, L.(2016) Acta Crystallogr F Struct Biol Commun 72: 863-869
- PubMed: 27917833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16017957
- Primary Citation Related Structures:
3UID - PubMed Abstract:
The structure of Msmeg_6760, a protein of unknown function, has been determined. Biochemical and bioinformatics analyses determined that Msmeg_6760 interacts with a protein encoded in the same operon, Msmeg_6762, and predicted that the operon is a toxin-antitoxin (TA) system. Structural comparison of Msmeg_6760 with proteins of known function suggests that Msmeg_6760 binds a hydrophobic ligand in a buried cavity lined by large hydrophobic residues. Access to this cavity could be controlled by a gate-latch mechanism. The function of the Msmeg_6760 toxin is unknown, but structure-based predictions revealed that Msmeg_6760 and Msmeg_6762 are homologous to Rv2034 and Rv2035, a predicted novel TA system involved in Mycobacterium tuberculosis latency during macrophage infection. The Msmeg_6760 toxin fold has not been previously described for bacterial toxins and its unique structural features suggest that toxin activation is likely to be mediated by a novel mechanism.
- UCLA-DOE Institute and Departments of Biological Chemistry and Chemistry and Biochemistry, University of California, Los Angeles, CA 90095-1570, USA.
Organizational Affiliation: