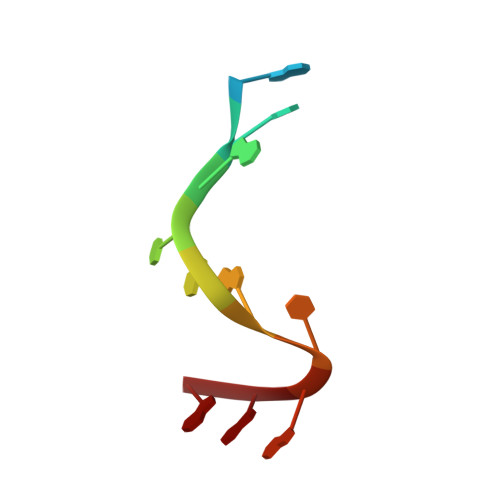
Structural basis for promoter-10 element recognition by the bacterial RNA polymerase sigma subunit.
Feklistov, A., Darst, S.A.(2011) Cell 147: 1257-1269
- PubMed: 22136875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2011.10.041
- Primary Citation Related Structures:
3UGO, 3UGP - PubMed Abstract:
The key step in bacterial promoter opening is recognition of the -10 promoter element (T(-12)A(-11)T(-10)A(-9)A(-8)T(-7) consensus sequence) by the RNA polymerase σ subunit. We determined crystal structures of σ domain 2 bound to single-stranded DNA bearing-10 element sequences. Extensive interactions occur between the protein and the DNA backbone of every -10 element nucleotide. Base-specific interactions occur primarily with A(-11) and T(-7), which are flipped out of the single-stranded DNA base stack and buried deep in protein pockets. The structures, along with biochemical data, support a model where the recognition of the -10 element sequence drives initial promoter opening as the bases of the nontemplate strand are extruded from the DNA double-helix and captured by σ. These results provide a detailed structural basis for the critical roles of A(-11) and T(-7) in promoter melting and reveal important insights into the initiation of transcription bubble formation.
- The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA. afeklistov@rockefeller.edu
Organizational Affiliation: