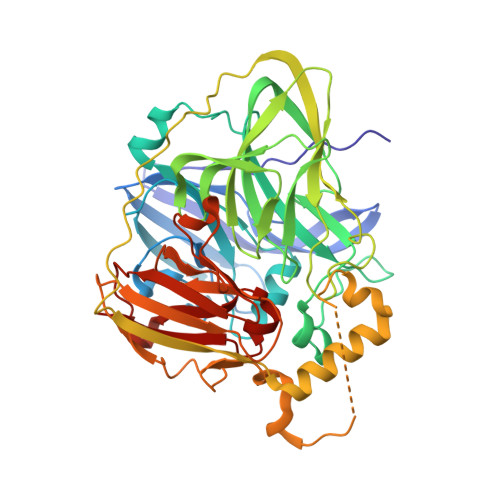
An O-centered structure of the trinuclear copper center in the Cys500Ser/Glu506Gln mutant of CueO and structural changes in low to high X-ray dose conditions.
Komori, H., Sugiyama, R., Kataoka, K., Higuchi, Y., Sakurai, T.(2012) Angew Chem Int Ed Engl 51: 1861-1864
- PubMed: 22250058 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201107739
- Primary Citation Related Structures:
3UAA, 3UAB, 3UAC, 3UAD, 3UAE - Graduate School of Life Science, University of Hyogo, 3-2-1, Koto, Kamigori-cho, Ako-gun 678-1297, Japan.
Organizational Affiliation: