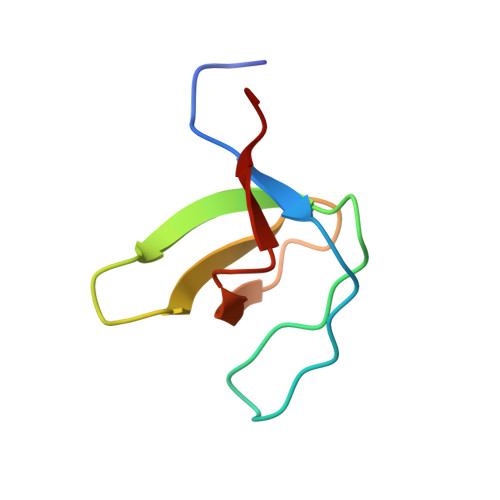
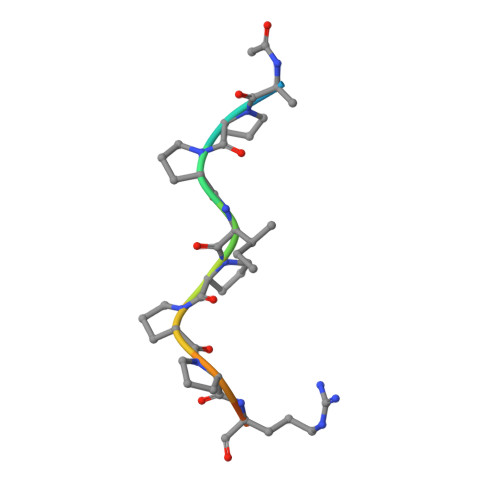
The promiscuous binding of the Fyn SH3 domain to a peptide from the NS5A protein.
Martin-Garcia, J.M., Luque, I., Ruiz-Sanz, J., Camara-Artigas, A.(2012) Acta Crystallogr D Biol Crystallogr 68: 1030-1040
- PubMed: 22868769 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912019798
- Primary Citation Related Structures:
3UA6, 3UA7 - PubMed Abstract:
The hepatitis C virus nonstructural 5A (NS5A) protein is a large zinc-binding phosphoprotein that plays an important role in viral RNA replication and is involved in altering signal transduction pathways in the host cell. This protein interacts with Fyn tyrosine kinase in vivo and regulates its kinase activity. The 1.5 Å resolution crystal structure of a complex between the SH3 domain of the Fyn tyrosine kinase and the C-terminal proline-rich motif of the NS5A-derived peptide APPIPPPRRKR has been solved. Crystals were obtained in the presence of ZnCl(2) and belonged to the tetragonal space group P4(1)2(1)2. The asymmetric unit is composed of four SH3 domains and two NS5A peptide molecules; only three of the domain molecules contain a bound peptide, while the fourth molecule seems to correspond to a free form of the domain. Additionally, two of the SH3 domains are bound to the same peptide chain and form a ternary complex. The proline-rich motif present in the NS5A protein seems to be important for RNA replication and virus assembly, and the promiscuous interaction of the Fyn SH3 domain with the NS5A C-terminal proline-rich peptide found in this crystallographic structure may be important in the virus infection cycle.
- Department of Physical Chemistry, Biochemistry and Inorganic Chemistry, University of Almería, Agrifood Campus of International Excellence (ceiA3), Carretera de Sacramento, 04120 Almería, Spain.
Organizational Affiliation: