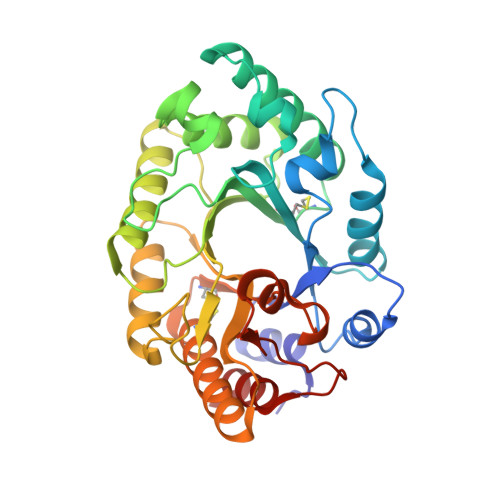
The structure of a GH10 xylanase from Fusarium oxysporum reveals the presence of an extended loop on top of the catalytic cleft.
Dimarogona, M., Topakas, E., Christakopoulos, P., Chrysina, E.D.(2012) Acta Crystallogr D Biol Crystallogr 68: 735-742
- PubMed: 22751658 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912007044
- Primary Citation Related Structures:
3U7B - PubMed Abstract:
Xylanase enzymes have been the focus of considerable research in recent decades owing to their extensive use in a variety of biotechnological applications. Previous structural studies of a number of GH10 xylanases revealed that all GH10 family members have the (β/α)(8)-barrel fold and their catalytic site is conserved. The structure of a new GH10 xylanase from Fusarium oxysporum (FoXyn10a) was determined at 1.94 Å resolution from crystals belonging to the tetragonal space group P4(1)2(1)2 with five molecules per asymmetric unit. Comparison of the structure of FoXyn10a with previously determined structures of GH10 family members indicated that most of the differences were located in the loop regions between the ordered secondary-structure elements of the barrel, as expected. However, alignment of FoXyn10a with sequence and structural homologues denoted an atypically long loop connecting strand β6b and helix α6 that was only present in one other GH10 xylanase, the structure of which is not known. This structural feature may be of functional importance, with potential implications in the catalytic efficiency of the enzyme.
- Institute of Organic and Pharmaceutical Chemistry, National Hellenic Research Foundation, 48 Vassileos Constantinou Avenue, 11635 Athens, Greece.
Organizational Affiliation: