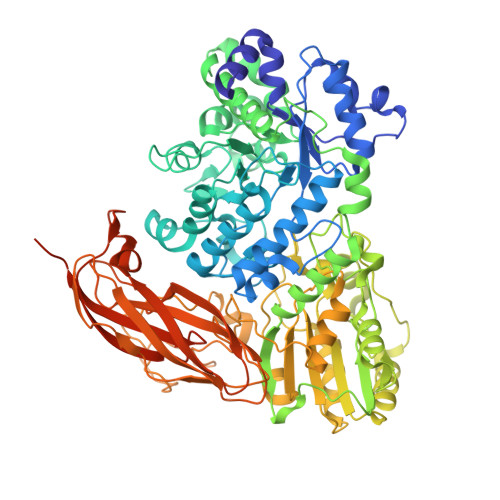
From soil to structure, a novel dimeric beta-glucosidase belonging to glycoside hydrolase family 3 isolated from compost using metagenomic analysis.
McAndrew, R.P., Park, J.I., Heins, R.A., Reindl, W., Friedland, G.D., D'haeseleer, P., Northen, T., Sale, K.L., Simmons, B.A., Adams, P.D.(2013) J Biological Chem 288: 14985-14992
- PubMed: 23580647 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.458356
- Primary Citation Related Structures:
3U48, 3U4A - PubMed Abstract:
A recent metagenomic analysis sequenced a switchgrass-adapted compost community to identify enzymes from microorganisms that were specifically adapted to switchgrass under thermophilic conditions. These enzymes are being examined as part of the pretreatment process for the production of "second-generation" biofuels. Among the enzymes discovered was JMB19063, a novel three-domain β-glucosidase that belongs to the GH3 (glycoside hydrolase 3) family. Here, we report the structure of JMB19063 in complex with glucose and the catalytic variant D261N crystallized in the presence of cellopentaose. JMB19063 is first structure of a dimeric member of the GH3 family, and we demonstrate that dimerization is required for catalytic activity. Arg-587 and Phe-598 from the C-terminal domain of the opposing monomer are shown to interact with bound ligands in the D261N structure. Enzyme assays confirmed that these residues are absolutely essential for full catalytic activity.
- Joint BioEnergy Institute, Emeryville, California 94608, USA.
Organizational Affiliation: