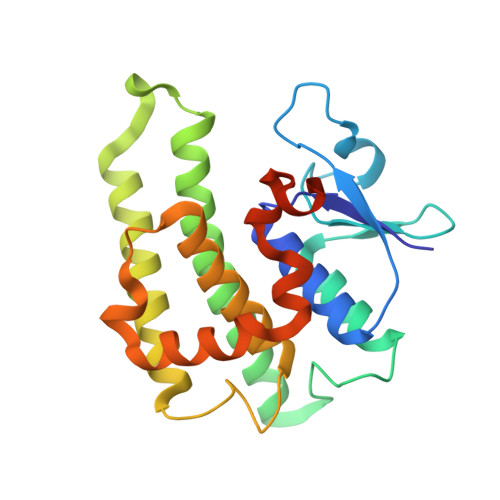
Crystal structure of GLUTATHIONE S-TRANSFERASE from Ralstonia solanacearum
Patskovsky, Y., Toro, R., Bhosle, R., Zencheck, W.D., Hillerich, B., Washington, R.D., Scott Glenn, A., Chowdhury, S., Evans, B., Hammonds, J., Imker, H.J., Armstrong, R.N., Gerlt, J.A., Almo, S.C.To be published.