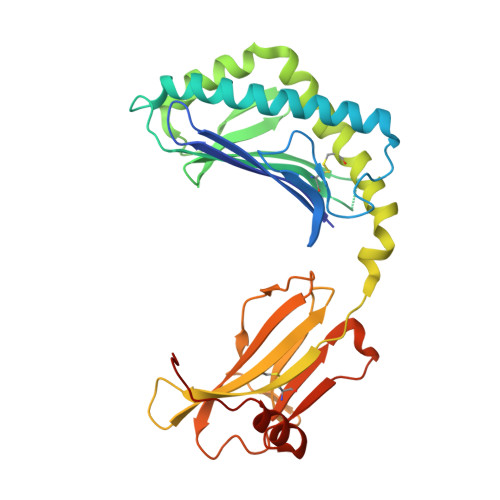
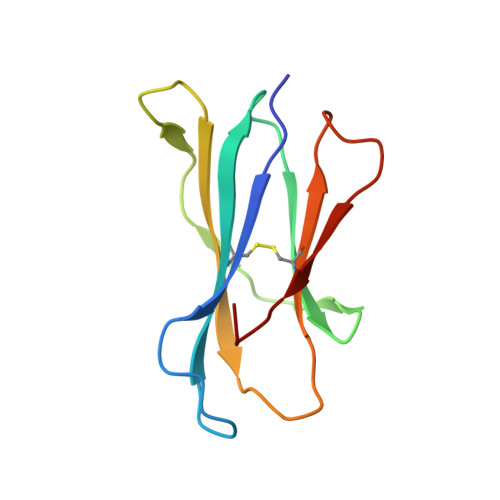
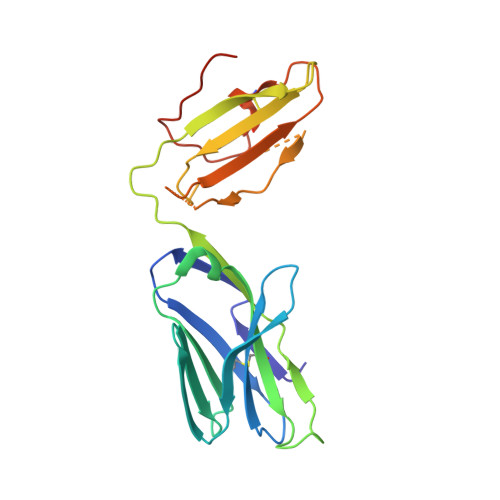
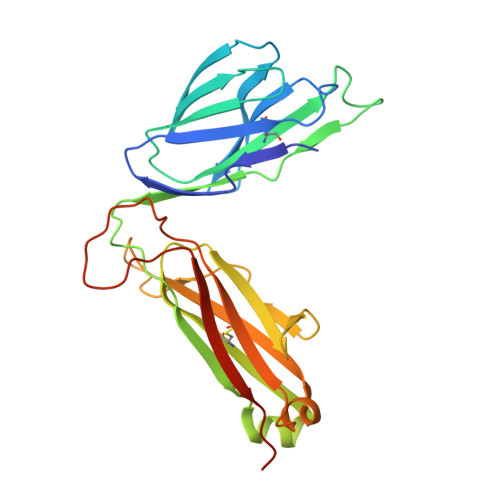
Vbeta2 natural killer T cell antigen receptor-mediated recognition of CD1d-glycolipid antigen.
Patel, O., Pellicci, D.G., Uldrich, A.P., Sullivan, L.C., Bhati, M., McKnight, M., Richardson, S.K., Howell, A.R., Mallevaey, T., Zhang, J., Bedel, R., Besra, G.S., Brooks, A.G., Kjer-Nielsen, L., McCluskey, J., Porcelli, S.A., Gapin, L., Rossjohn, J., Godfrey, D.I.(2011) Proc Natl Acad Sci U S A 108: 19007-19012
- PubMed: 22065767 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1109066108
- Primary Citation Related Structures:
3TO4 - PubMed Abstract:
Natural killer T cell antigen receptors (NKT TCRs) recognize lipid-based antigens (Ags) presented by CD1d. Although the TCR α-chain is invariant, NKT TCR Vβ exhibits greater diversity, with one (Vβ11) and three (Vβ8, Vβ7, and Vβ2) Vβ chains in humans and mice, respectively. With the exception of the Vβ2 NKT TCR, NKT TCRs possess canonical tyrosine residues within complementarity determining region (CDR) 2β that are critical for CD1d binding. Thus, how Vβ2 NKT TCR docks with CD1d-Ag was unclear. Despite the absence of the CDR2β-encoded tyrosine residues, we show that the Vβ2 NKT TCR engaged CD1d-Ag in a similar manner and with a comparable affinity and energetic footprint to the manner observed for the Vβ8.2 and Vβ7 NKT TCRs. Accordingly, the germline-encoded regions of the TCR β-chain do not exclusively dictate the innate NKT TCR-CD1d-Ag docking mode. Nevertheless, clear fine specificity differences for the CD1d-Ag existed between the Vβ2 NKT TCR and the Vβ8.2 and Vβ7 NKT TCRs, with the Vβ2 NKT TCR exhibiting greater sensitivity to modifications to the glycolipid Ag. Furthermore, within the Vβ2 NKT TCR-CD1d-αGalCer complex, the CDR2β loop mediated fewer contacts with CD1d, whereas the CDR1β and CDR3β loops contacted CD1d to a much greater extent compared with most Vβ11, Vβ8.2, and Vβ7 NKT TCRs. Accordingly, there is a greater interplay between the germline- and nongermline-encoded loops within the TCR β-chain of the Vβ2 NKT TCR that enables CD1d-Ag ligation.
- Protein Crystallography Unit, Australian Research Council Centre of Excellence in Structural and Functional Microbial Genomics, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: