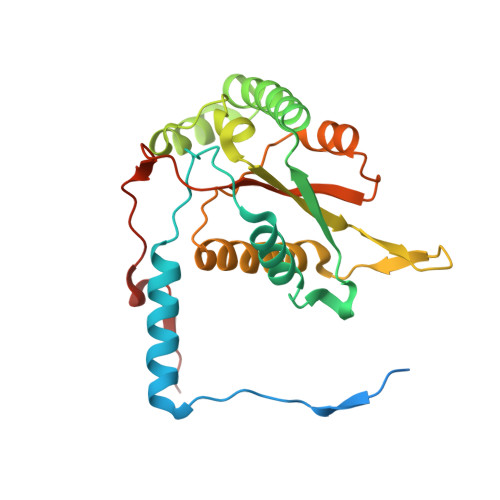
Expression of a soluble form of iodotyrosine deiodinase for active site characterization by engineering the native membrane protein from Mus musculus.
Buss, J.M., McTamney, P.M., Rokita, S.E.(2012) Protein Sci 21: 351-361
- PubMed: 22238141 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2020
- Primary Citation Related Structures:
3TNZ, 3TO0 - PubMed Abstract:
Reductive deiodination is critical for thyroid function and represents an unusual exception to the more common oxidative and hydrolytic mechanisms of dehalogenation in mammals. Studies on the reductive processes have been limited by a lack of convenient methods for heterologous expression of the appropriate proteins in large scale. The enzyme responsible for iodide salvage in the thyroid, iodotyrosine deodinase, is now readily generated after engineering its gene from Mus musculus. High expression of a truncated derivative lacking the membrane domain at its N-terminal was observed in Sf9 cells, whereas expression in Pichia pastoris remained low despite codon optimization. Ultimately, the desired expression in Escherichia coli was achieved after replacing the two conserved Cys residues of the deiodinase with Ala and fusing the resulting protein to thioredoxin. This final construct provided abundant enzyme for crystallography and mutagenesis. Utility of the E. coli system was demonstrated by examining a set of active site residues critical for binding to the zwitterionic portion of substrate.
- Department of Chemistry and Biochemistry, University of Maryland, College Park, Maryland 20742-2021, USA.
Organizational Affiliation: