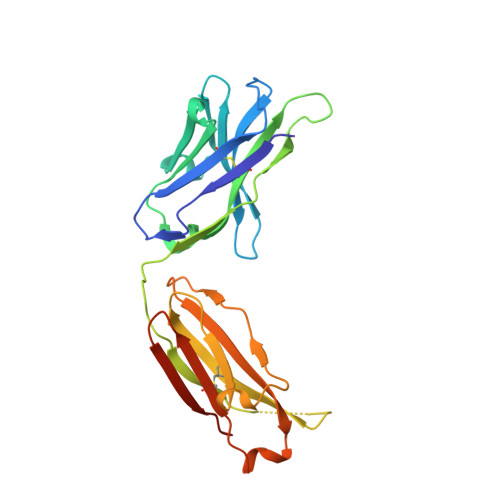
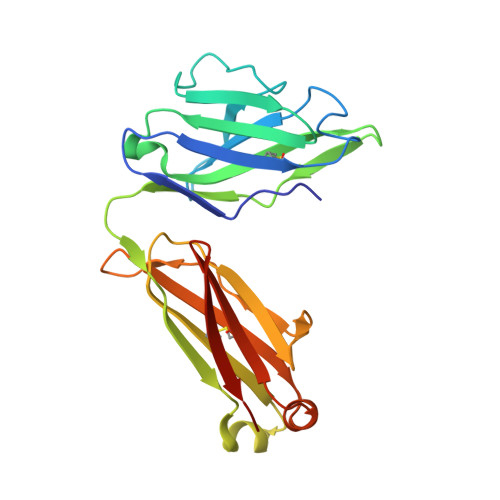
Diverse specificity and effector function among human antibodies to HIV-1 envelope glycoprotein epitopes exposed by CD4 binding.
Guan, Y., Pazgier, M., Sajadi, M.M., Kamin-Lewis, R., Al-Darmarki, S., Flinko, R., Lovo, E., Wu, X., Robinson, J.E., Seaman, M.S., Fouts, T.R., Gallo, R.C., DeVico, A.L., Lewis, G.K.(2013) Proc Natl Acad Sci U S A 110: E69-E78
- PubMed: 23237851 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1217609110
- Primary Citation Related Structures:
3TNM - PubMed Abstract:
The HIV-1 envelope glycoprotein (Env) undergoes conformational transitions consequent to CD4 binding and coreceptor engagement during viral entry. The physical steps in this process are becoming defined, but less is known about their significance as targets of antibodies potentially protective against HIV-1 infection. Here we probe the functional significance of transitional epitope exposure by characterizing 41 human mAbs specific for epitopes exposed on trimeric Env after CD4 engagement. These mAbs recognize three epitope clusters: cluster A, the gp120 face occluded by gp41 in trimeric Env; cluster B, a region proximal to the coreceptor-binding site (CoRBS) and involving the V1/V2 domain; and cluster C, the coreceptor-binding site. The mAbs were evaluated functionally by antibody-dependent, cell-mediated cytotoxicity (ADCC) and for neutralization of Tiers 1 and 2 pseudoviruses. All three clusters included mAbs mediating ADCC. However, there was a strong potency bias for cluster A, which harbors at least three potent ADCC epitopes whose cognate mAbs have electropositive paratopes. Cluster A epitopes are functional ADCC targets during viral entry in an assay format using virion-sensitized target cells. In contrast, only cluster C contained epitopes that were recognized by neutralizing mAbs. There was significant diversity in breadth and potency that correlated with epitope fine specificity. In contrast, ADCC potency had no relationship with neutralization potency or breadth for any epitope cluster. Thus, Fc-mediated effector function and neutralization coselect with specificity in anti-Env antibody responses, but the nature of selection is distinct for these two antiviral activities.
- Division of Basic Science and Vaccine Research, Institute of Human Virology, University of Maryland School of Medicine, Baltimore, MD 21201, USA.
Organizational Affiliation: