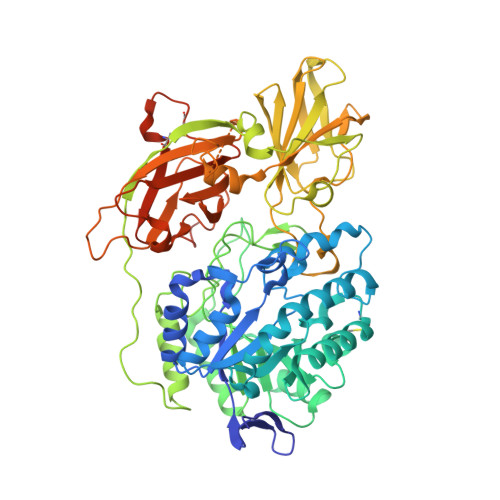
Crystal structure of human beta-galactosidase: structural basis of Gm1 gangliosidosis and morquio B diseases
Ohto, U., Usui, K., Ochi, T., Yuki, K., Satow, Y., Shimizu, T.(2012) J Biological Chem 287: 1801-1812
- PubMed: 22128166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.293795
- Primary Citation Related Structures:
3THC, 3THD - PubMed Abstract:
G(M1) gangliosidosis and Morquio B are autosomal recessive lysosomal storage diseases associated with a neurodegenerative disorder or dwarfism and skeletal abnormalities, respectively. These diseases are caused by deficiencies in the lysosomal enzyme β-d-galactosidase (β-Gal), which lead to accumulations of the β-Gal substrates, G(M1) ganglioside, and keratan sulfate. β-Gal is an exoglycosidase that catalyzes the hydrolysis of terminal β-linked galactose residues. This study shows the crystal structures of human β-Gal in complex with its catalytic product galactose or with its inhibitor 1-deoxygalactonojirimycin. Human β-Gal is composed of a catalytic TIM barrel domain followed by β-domain 1 and β-domain 2. To gain structural insight into the molecular defects of β-Gal in the above diseases, the disease-causing mutations were mapped onto the three-dimensional structure. Finally, the possible causes of the diseases are discussed.
- From the Graduate School of Pharmaceutical Sciences, University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: