T cell receptor signaling is limited by docking geometry to peptide-major histocompatibility complex.
Adams, J.J., Narayanan, S., Liu, B., Birnbaum, M.E., Kruse, A.C., Bowerman, N.A., Chen, W., Levin, A.M., Connolly, J.M., Zhu, C., Kranz, D.M., Garcia, K.C.(2011) Immunity 35: 681-693
- PubMed: 22101157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2011.09.013
- Primary Citation Related Structures:
3TF7, 3TFK, 3TJH, 3TPU - PubMed Abstract:
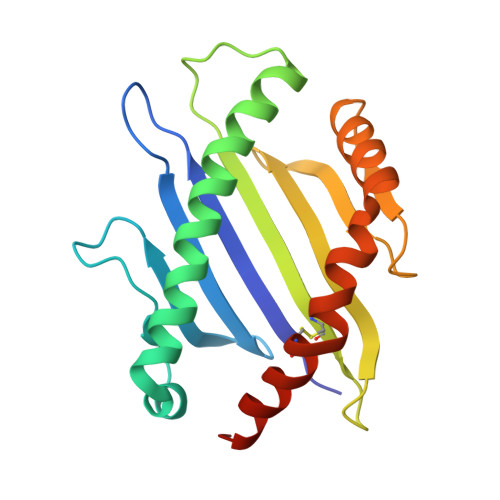
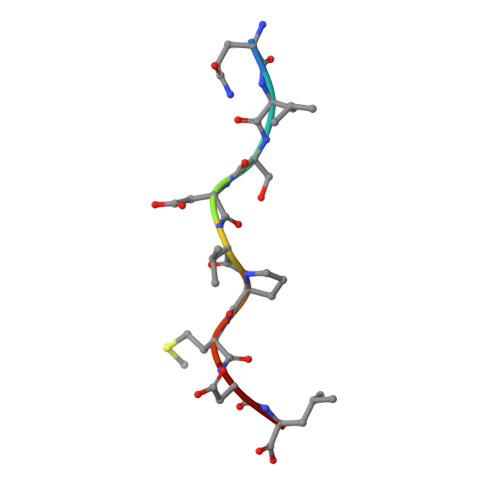
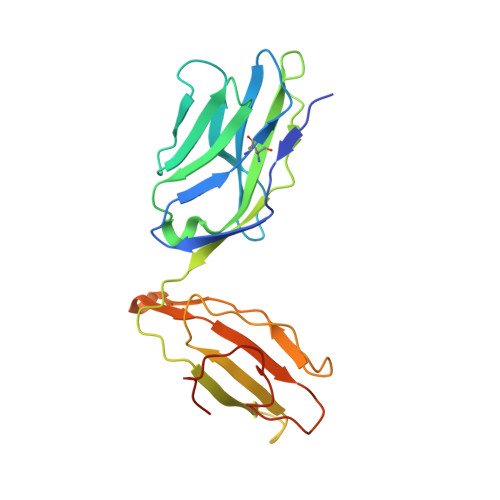
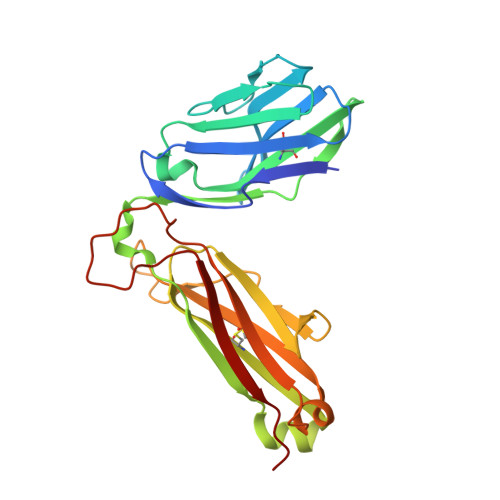
T cell receptor (TCR) engagement of peptide-major histocompatibility complex (pMHC) is essential to adaptive immunity, but it is unknown whether TCR signaling responses are influenced by the binding topology of the TCR-peptide-MHC complex. We developed yeast-displayed pMHC libraries that enabled us to identify new peptide sequences reactive with a single TCR. Structural analysis showed that four peptides bound to the TCR with distinct 3D and 2D affinities using entirely different binding chemistries. Three of the peptides that shared a common docking mode, where key TCR-MHC germline interactions are preserved, induced TCR signaling. The fourth peptide failed to induce signaling and was recognized in a substantially different TCR-MHC binding mode that apparently exceeded geometric tolerances compatible with signaling. We suggest that the stereotypical TCR-MHC docking paradigm evolved from productive signaling geometries and that TCR signaling can be modulated by peptides that are recognized in alternative TCR-pMHC binding orientations.
- Howard Hughes Medical Institute, and Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: