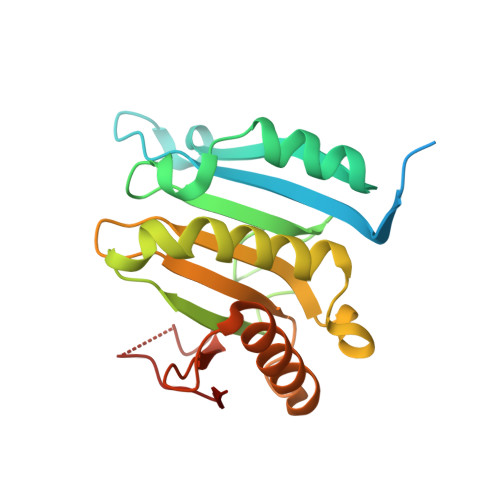
Structural insights into the allosteric effects of 4EBP1 on the eukaryotic translation initiation factor eIF4E.
Siddiqui, N., Tempel, W., Nedyalkova, L., Volpon, L., Wernimont, A.K., Osborne, M.J., Park, H.W., Borden, K.L.(2012) J Mol Biology 415: 781-792
- PubMed: 22178476 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.12.002
- Primary Citation Related Structures:
3TF2, 3U7X - PubMed Abstract:
The eukaryotic translation initiation factor eIF4E plays key roles in cap-dependent translation and mRNA export. These functions rely on binding the 7-methyl-guanosine moiety (5'cap) on the 5'-end of all mRNAs. eIF4E is regulated by proteins such as eIF4G and eIF4E binding proteins (4EBPs) that bind the dorsal surface of eIF4E, distal to the cap binding site, and modulate cap binding activity. Both proteins increase the affinity of eIF4E for 5'cap. Our understanding of the allosteric effects and structural underpinnings of 4EBP1 or eIF4G binding can be advanced by obtaining structural data on cap-free eIF4E bound to one of these proteins. Here, we report the crystal structure of apo-eIF4E and cap-free eIF4E in complex with a 4EBP1 peptide. We also monitored 4EBP1 binding to cap-free eIF4E in solution using NMR. Together, these studies suggest that 4EBP1 transforms eIF4E into a cap-receptive state. NMR methods were also used to compare the allosteric routes activated by 4EBP1, eIF4G, and the arenavirus Z protein, a negative regulator of cap binding. We observed chemical shift perturbation at the dorsal binding site leading to alterations in the core of the protein, which were ultimately communicated to the unoccupied cap binding site of eIF4E. There were notable similarities between the routes taken by 4EBP1 and eIF4G and differences from the negative regulator Z. Thus, binding of 4EBP1 or eIF4G allosterically drives alterations throughout the protein that increase the affinity of eIF4E for the 5'cap.
- Institute for Research in Immunology and Cancer, Université de Montréal, Montréal, QC, Canada H3T 1J4.
Organizational Affiliation: