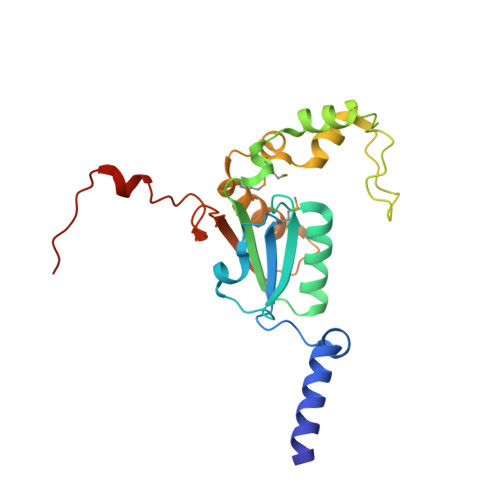
Evolution of a new enzyme for carbon disulphide conversion by an acidothermophilic archaeon.
Smeulders, M.J., Barends, T.R., Pol, A., Scherer, A., Zandvoort, M.H., Udvarhelyi, A., Khadem, A.F., Menzel, A., Hermans, J., Shoeman, R.L., Wessels, H.J., van den Heuvel, L.P., Russ, L., Schlichting, I., Jetten, M.S., Op den Camp, H.J.(2011) Nature 478: 412-416
- PubMed: 22012399 Search on PubMed
- DOI: https://doi.org/10.1038/nature10464
- Primary Citation Related Structures:
3TEN, 3TEO - PubMed Abstract:
Extremophilic organisms require specialized enzymes for their exotic metabolisms. Acid-loving thermophilic Archaea that live in the mudpots of volcanic solfataras obtain their energy from reduced sulphur compounds such as hydrogen sulphide (H(2)S) and carbon disulphide (CS(2)). The oxidation of these compounds into sulphuric acid creates the extremely acidic environment that characterizes solfataras. The hyperthermophilic Acidianus strain A1-3, which was isolated from the fumarolic, ancient sauna building at the Solfatara volcano (Naples, Italy), was shown to rapidly convert CS(2) into H(2)S and carbon dioxide (CO(2)), but nothing has been known about the modes of action and the evolution of the enzyme(s) involved. Here we describe the structure, the proposed mechanism and evolution of a CS(2) hydrolase from Acidianus A1-3. The enzyme monomer displays a typical β-carbonic anhydrase fold and active site, yet CO(2) is not one of its substrates. Owing to large carboxy- and amino-terminal arms, an unusual hexadecameric catenane oligomer has evolved. This structure results in the blocking of the entrance to the active site that is found in canonical β-carbonic anhydrases and the formation of a single 15-Å-long, highly hydrophobic tunnel that functions as a specificity filter. The tunnel determines the enzyme's substrate specificity for CS(2), which is hydrophobic. The transposon sequences that surround the gene encoding this CS(2) hydrolase point to horizontal gene transfer as a mechanism for its acquisition during evolution. Our results show how the ancient β-carbonic anhydrase, which is central to global carbon metabolism, was transformed by divergent evolution into a crucial enzyme in CS(2) metabolism.
- Department of Microbiology, Radboud University Nijmegen, Heyendaalseweg 135, 6525 AJ, Nijmegen, The Netherlands.
Organizational Affiliation: