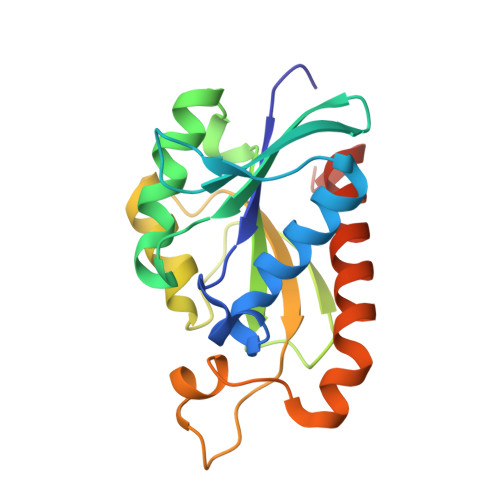
Structures of new crystal forms of Mycobacterium tuberculosis peptidyl-tRNA hydrolase and functionally important plasticity of the molecule
Selvaraj, M., Ahmad, R., Varshney, U., Vijayan, M.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 124-128
- PubMed: 22297982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111052341
- Primary Citation Related Structures:
3TCK, 3TCN, 3TD2, 3TD6 - PubMed Abstract:
The X-ray structures of new crystal forms of peptidyl-tRNA hydrolase from M. tuberculosis reported here and the results of previous X-ray studies of the enzyme from different sources provide a picture of the functionally relevant plasticity of the protein molecule. The new X-ray results confirm the connection deduced previously between the closure of the lid at the peptide-binding site and the opening of the gate that separates the peptide-binding and tRNA-binding sites. The plasticity of the molecule indicated by X-ray structures is in general agreement with that deduced from the available solution NMR results. The correlation between the lid and the gate movements is not, however, observed in the NMR structure.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: