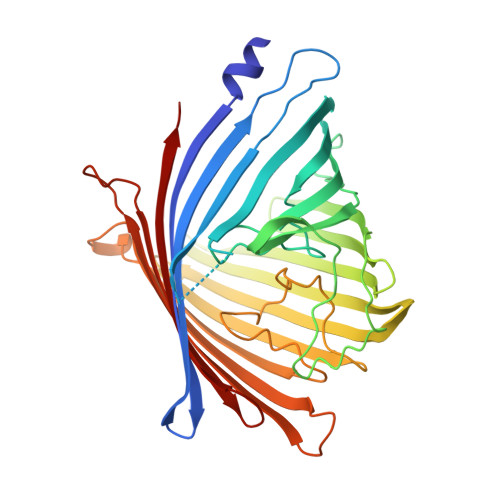
Substrate Specificity within a Family of Outer Membrane Carboxylate Channels.
Eren, E., Vijayaraghavan, J., Liu, J., Cheneke, B.R., Touw, D.S., Lepore, B.W., Indic, M., Movileanu, L., van den Berg, B.(2012) PLoS Biol 10: e1001242-e1001242
- PubMed: 22272184 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1001242
- Primary Citation Related Structures:
3SY7, 3SY9, 3SYB, 3SYS, 3SZD, 3SZV, 3T0S, 3T20, 3T24 - PubMed Abstract:
Many Gram-negative bacteria, including human pathogens such as Pseudomonas aeruginosa, do not have large-channel porins. This results in an outer membrane (OM) that is highly impermeable to small polar molecules, making the bacteria intrinsically resistant towards many antibiotics. In such microorganisms, the majority of small molecules are taken up by members of the OprD outer membrane protein family. Here we show that OprD channels require a carboxyl group in the substrate for efficient transport, and based on this we have renamed the family Occ, for outer membrane carboxylate channels. We further show that Occ channels can be divided into two subfamilies, based on their very different substrate specificities. Our results rationalize how certain bacteria can efficiently take up a variety of substrates under nutrient-poor conditions without compromising membrane permeability. In addition, they explain how channel inactivation in response to antibiotics can cause resistance but does not lead to decreased fitness.
- University of Massachusetts Medical School, Program in Molecular Medicine, Worcester, Massachusetts, United States of America.
Organizational Affiliation: