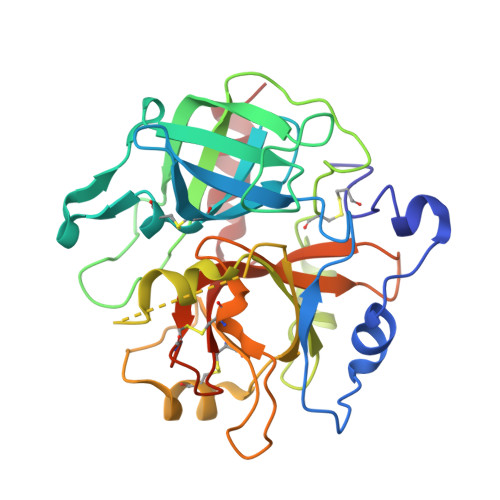
Crystal structures of prethrombin-2 reveal alternative conformations under identical solution conditions and the mechanism of zymogen activation.
Pozzi, N., Chen, Z., Zapata, F., Pelc, L.A., Barranco-Medina, S., Di Cera, E.(2011) Biochemistry 50: 10195-10202
- PubMed: 22049947 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi2015019
- Primary Citation Related Structures:
3SQE, 3SQH - PubMed Abstract:
Prethrombin-2 is the immediate zymogen precursor of the clotting enzyme thrombin, which is generated upon cleavage at R15 and separation of the A chain and catalytic B chain. The X-ray structure of prethrombin-2 determined in the free form at 1.9 Å resolution shows the 215-217 segment collapsed into the active site and occluding 49% of the volume available for substrate binding. Remarkably, some of the crystals harvested from the same crystallization well, under identical solution conditions, diffract to 2.2 Å resolution in the same space group but produce a structure in which the 215-217 segment moves >5 Å and occludes 24% of the volume available for substrate binding. The two alternative conformations of prethrombin-2 have the side chain of W215 relocating >9 Å within the active site and are relevant to the allosteric E*-E equilibrium of the mature enzyme. Another unanticipated feature of prethrombin-2 bears on the mechanism of prothrombin activation. R15 is found buried within the protein in ionic interactions with E14e, D14l, and E18, thereby making its exposure to solvent necessary for proteolytic attack and conversion to thrombin. On the basis of this structural observation, we constructed the E14eA/D14lA/E18A triple mutant to reduce the level of electrostatic coupling with R15 and promote zymogen activation. The mutation causes prethrombin-2 to spontaneously convert to thrombin, without the need for the snake venom ecarin or the physiological prothrombinase complex.
- Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, Missouri 63104, United States.
Organizational Affiliation: