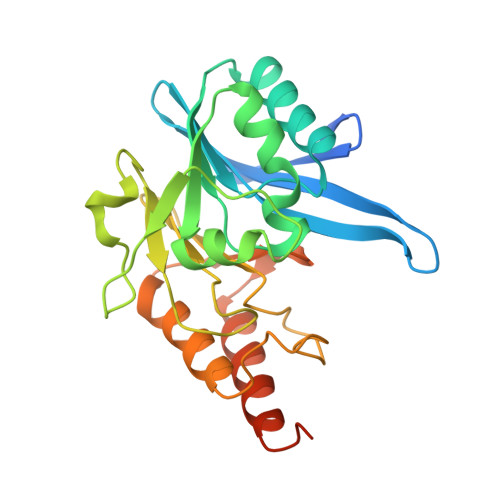
Crystal structure of New Delhi metallo-beta-lactamase reveals molecular basis for antibiotic resistance
King, D., Strynadka, N.(2011) Protein Sci 20: 1484-1491
- PubMed: 21774017 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.697
- Primary Citation Related Structures:
3SPU - PubMed Abstract:
β-Lactams are the most commonly prescribed class of antibiotics and have had an enormous impact on human health. Thus, it is disquieting that an enzyme called New Delhi metallo-β-lactamase-1 (NDM-1) can confer Enterobacteriaceae with nearly complete resistance to all β-lactam antibiotics including the carbapenams. We have determined the crystal structure of Klebsiella pneumoniae apo-NDM-1 to 2.1-Å resolution. From the structure, we see that NDM-1 has an expansive active site with a unique electrostatic profile, which we propose leads to a broader substrate specificity. In addition, NDM-1 undergoes important conformational changes upon substrate binding. These changes have not been previously observed in metallo-β-lactamase enzymes and may have a direct influence on substrate recognition and catalysis.
- Department of Biochemistry and Molecular Biology and Center for Blood Research, University of British Columbia, Vancouver, British Columbia, Canada.
Organizational Affiliation: