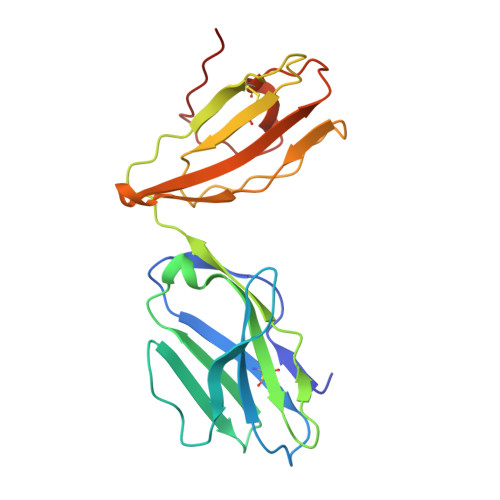
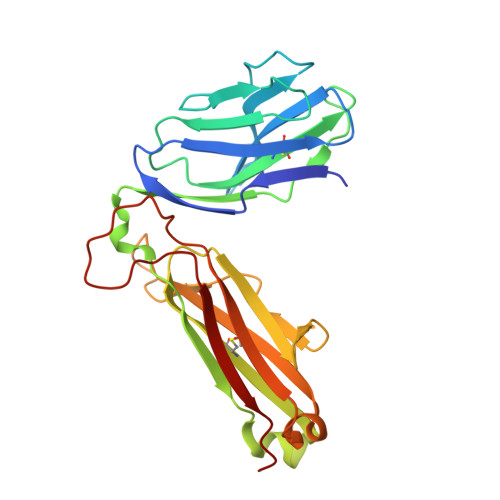
A structural basis for varied alpha-beta TCR usage against an immunodominant EBV antigen restricted to a HLA-B8 molecule.
Gras, S., Wilmann, P.G., Chen, Z., Halim, H., Liu, Y.C., Kjer-Nielsen, L., Purcell, A.W., Burrows, S.R., McCluskey, J., Rossjohn, J.(2012) J Immunol 188: 311-321
- PubMed: 22140258 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.1102686
- Primary Citation Related Structures:
3SJV, 3SKM, 3SKN, 3SKO - PubMed Abstract:
EBV is a ubiquitous and persistent human pathogen, kept in check by the cytotoxic T cell response. In this study, we investigated how three TCRs, which differ in their T cell immunodominance hierarchies and gene usage, interact with the same EBV determinant (FLRGRAYGL), bound to the same Ag-presenting molecule, HLA-B8. We found that the three TCRs exhibit differing fine specificities for the viral Ag. Further, via structural and biophysical approaches, we demonstrated that the viral Ag provides the greatest energetic contribution to the TCR-peptide-HLA interaction, while focusing on a few adjacent HLA-based interactions to further tune fine-specificity requirements. Thus, the TCR engages the peptide-HLA with the viral Ag as the main glue, such that neighboring TCR-MHC interactions are recruited as a supportive adhesive. Collectively, we provide a portrait of how the host's adaptive immune response differentially engages a common viral Ag.
- Department of Biochemistry and Molecular Biology, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: