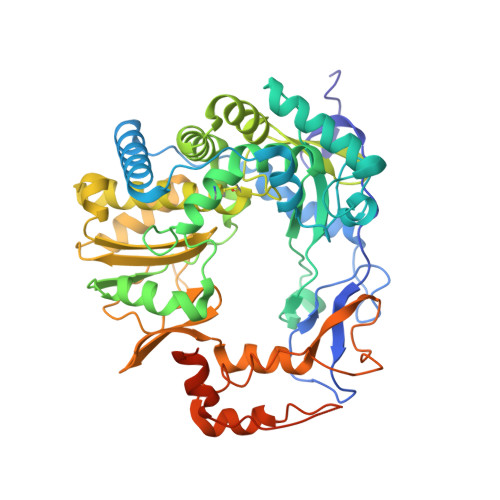
Crystal structures of murine norovirus-1 RNA-dependent RNA polymerase in complex with 2-thiouridine or ribavirin.
Alam, I., Lee, J.H., Cho, K.J., Han, K.R., Yang, J.M., Chung, M.S., Kim, K.H.(2012) Virology 426: 143-151
- PubMed: 22341781 Search on PubMed
- DOI: https://doi.org/10.1016/j.virol.2012.01.016
- Primary Citation Related Structures:
3SFG, 3SFU - PubMed Abstract:
Murine norovirus-1 (MNV-1) shares many features with human norovirus (HuNoV) and both are classified within the norovirus genus of Caliciviridae family. MNV-1 is used as the surrogate for HuNoV research since it is the only form that can be grown in cell culture. HuNoV and MNV-1 RNA dependent RNA polymerase (RdRp) proteins with the sequence identity of 59% show essentially identical conformations. Here we report the first structural evidence of 2-thiouridine (2TU) or ribavirin binding to MNV-1 RdRp, based on the crystal structures determined at 2.2Å and 2.5Å resolutions, respectively. Cellular and biochemical studies revealed stronger inhibitory effect of 2TU on the replication of MNV-1 in RAW 264.7 cells, compared to that of ribavirin. Our complex structures highlight the key interactions involved in recognition of the nucleoside analogs which block the active site of the viral RNA polymerase.
- Department of Biotechnology & Bioinformatics, Korea University, Chungnam 339-700, Republic of Korea.
Organizational Affiliation: