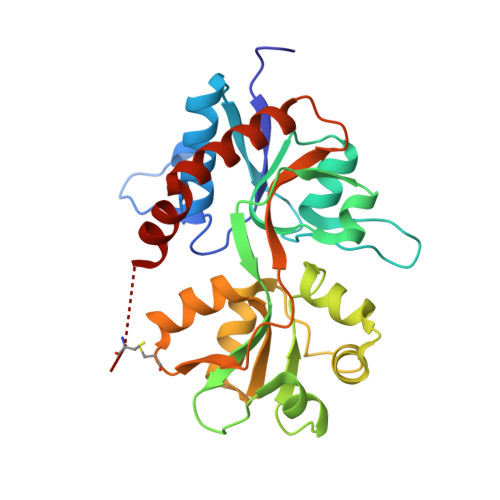
Binding site and interlobe interactions of the ionotropic glutamate receptor GluK3 ligand binding domain revealed by high resolution crystal structure in complex with (S)-glutamate.
Venskutonyte, R., Frydenvang, K., Gajhede, M., Bunch, L., Pickering, D.S., Kastrup, J.S.(2011) J Struct Biol 176: 307-314
- PubMed: 21907808 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2011.08.014
- Primary Citation Related Structures:
3S9E, 4MH5 - PubMed Abstract:
Ionotropic glutamate receptors (iGluRs) are involved in excitatory signal transmission throughout the central nervous system and their malfunction is associated with various health disorders. GluK3 is a subunit of iGluRs, belonging to the subfamily of kainate receptors (GluK1-5). Several crystal structures of GluK1 and GluK2 ligand binding domains have been determined in complex with agonists and antagonists. However, little is known about the molecular mechanisms underlying GluK3 ligand binding properties and no compounds displaying reasonable selectivity towards GluK3 are available today. Here, we present the first X-ray crystal structure of the ligand binding domain of GluK3 in complex with glutamate, determined to 1.6Å resolution. The structure reveals a conserved glutamate binding mode, characteristic for iGluRs, and a water molecule network in the glutamate binding site similar to that seen in GluK1. In GluK3, a slightly lower degree of domain closure around glutamate is observed compared to most other kainate receptor structures with glutamate. The volume of the GluK3 glutamate binding cavity was found to be of intermediate size between those of GluK1 and GluK2. The residues in GluK3 contributing to the subfamily differences in the binding sites are primarily: Thr520, Ala691, Asn722, Leu736 and Thr742. The GluK3 ligand binding domain seems to be less stabilized through interlobe interactions than GluK1 and this may contribute to the faster desensitization kinetics of GluK3.
- Department of Medicinal Chemistry, University of Copenhagen, Universitetsparken 2, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: