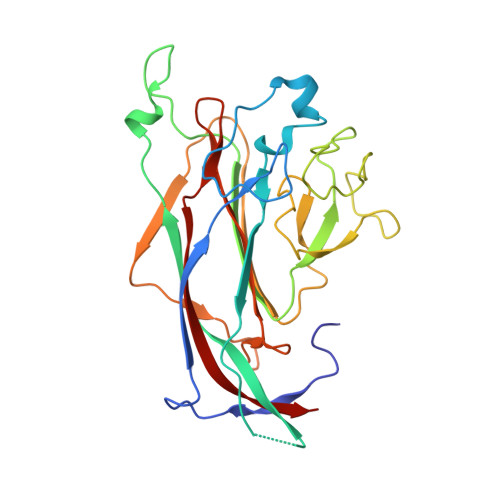
Structures of the major capsid proteins of the human KI and WU polyomaviruses.
Neu, U., Wang, J., Macejak, D., Garcea, R.L., Stehle, T.(2011) J Virol 85: 7384-7392
- PubMed: 21543504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00382-11
- Primary Citation Related Structures:
3S7V, 3S7X - PubMed Abstract:
The Karolinska Institutet and Washington University polyomaviruses (KIPyV and WUPyV, respectively) are recently discovered human viruses that infect the respiratory tract. Although they have not yet been linked to disease, they are prevalent in populations worldwide, with initial infection occurring in early childhood. Polyomavirus capsids consist of 72 pentamers of the major capsid protein viral protein 1 (VP1), which determines antigenicity and receptor specificity. The WUPyV and KIPyV VP1 proteins are distant in evolution from VP1 proteins of known structure such as simian virus 40 or murine polyomavirus. We present here the crystal structures of unassembled recombinant WUPyV and KIPyV VP1 pentamers at resolutions of 2.9 and 2.55 Å, respectively. The WUPyV and KIPyV VP1 core structures fold into the same β-sandwich that is a hallmark of all polyomavirus VP1 proteins crystallized to date. However, differences in sequence translate into profoundly different surface loop structures in KIPyV and WUPyV VP1 proteins. Such loop structures have not been observed for other polyomaviruses, and they provide initial clues about the possible interactions of these viruses with cell surface receptors.
- Interfaculty Institute of Biochemistry, University of Tuebingen, Hoppe-Seyler-Str. 4, D-72076 Tuebingen, Germany.
Organizational Affiliation: