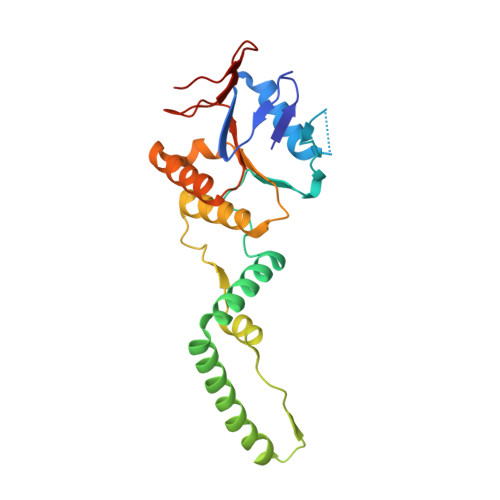
Crystal structure of clustered regularly interspaced short palindromic repeats (CRISPR)-associated Csn2 protein revealed Ca2+-dependent double-stranded DNA binding activity.
Nam, K.H., Kurinov, I., Ke, A.(2011) J Biological Chem 286: 30759-30768
- PubMed: 21697083 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.256263
- Primary Citation Related Structures:
3S5U - PubMed Abstract:
Clustered regularly interspaced short palindromic repeats (CRISPR) and their associated protein genes (cas genes) are widespread in bacteria and archaea. They form a line of RNA-based immunity to eradicate invading bacteriophages and malicious plasmids. A key molecular event during this process is the acquisition of new spacers into the CRISPR loci to guide the selective degradation of the matching foreign genetic elements. Csn2 is a Nmeni subtype-specific cas gene required for new spacer acquisition. Here we characterize the Enterococcus faecalis Csn2 protein as a double-stranded (ds-) DNA-binding protein and report its 2.7 Å tetrameric ring structure. The inner circle of the Csn2 tetrameric ring is ∼26 Å wide and populated with conserved lysine residues poised for nonspecific interactions with ds-DNA. Each Csn2 protomer contains an α/β domain and an α-helical domain; significant hinge motion was observed between these two domains. Ca(2+) was located at strategic positions in the oligomerization interface. We further showed that removal of Ca(2+) ions altered the oligomerization state of Csn2, which in turn severely decreased its affinity for ds-DNA. In summary, our results provided the first insight into the function of the Csn2 protein in CRISPR adaptation by revealing that it is a ds-DNA-binding protein functioning at the quaternary structure level and regulated by Ca(2+) ions.
- Department of Molecular Biology and Genetics, Cornell University, Ithaca, New York 14850.
Organizational Affiliation: