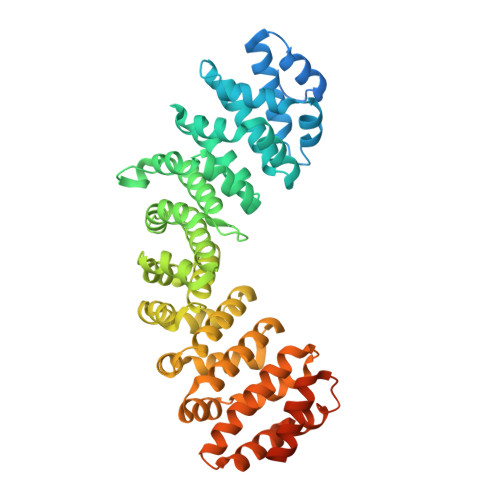
Structural basis of importin-alpha-mediated nuclear transport for Ku70 and Ku80.
Takeda, A.A., de Barros, A.C., Chang, C.W., Kobe, B., Fontes, M.R.(2011) J Mol Biology 412: 226-234
- PubMed: 21806995 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.07.038
- Primary Citation Related Structures:
3RZ9, 3RZX - PubMed Abstract:
Ku70 and Ku80 form a heterodimeric complex involved in multiple nuclear processes. This complex plays a key role in DNA repair due to its ability to bind DNA double-strand breaks and facilitate repair by the nonhomologous end-joining pathway. Ku70 and Ku80 have been proposed to contain bipartite and monopartite nuclear localization sequences (NLSs), respectively, that allow them to be translocated to the nucleus independently of each other via the classical importin-α (Impα)/importin-β-mediated nuclear import pathway. To determine the structural basis of the recognition of Ku70 and Ku80 proteins by Impα, we solved the crystal structures of the complexes of Impα with the peptides corresponding to the Ku70 and Ku80 NLSs. Our structural studies confirm the binding of the Ku80 NLS as a classical monopartite NLS but reveal an unexpected binding mode for Ku70 NLS with only one basic cluster bound to the receptor. Both Ku70 and Ku80 therefore contain monopartite NLSs, and sequences outside the basic cluster make favorable interactions with Impα, suggesting that this may be a general feature in monopartite NLSs. We show that the Ku70 NLS has a higher affinity for Impα than the Ku80 NLS, consistent with more extensive interactions in its N-terminal region. The prospect of nuclear import of Ku70 and Ku80 independently of each other provides a powerful regulatory mechanism for the function of the Ku70/Ku80 heterodimer and independent functions of the two proteins.
- Departamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista, Botucatu, SP 18618-970, Brazil.
Organizational Affiliation: