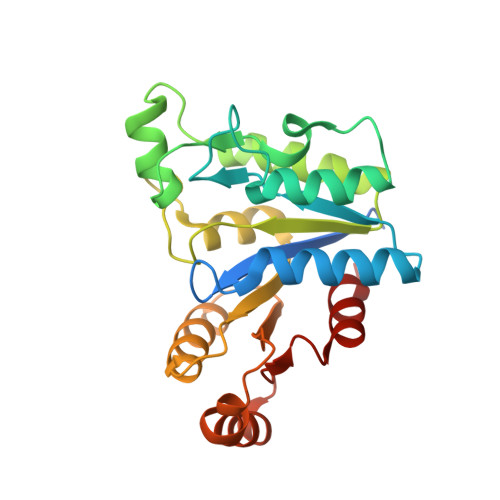
Structural characterization of Helicobacter pylori dethiobiotin synthetase reveals differences between family members.
Porebski, P.J., Klimecka, M., Chruszcz, M., Nicholls, R.A., Murzyn, K., Cuff, M.E., Xu, X., Cymborowski, M., Murshudov, G.N., Savchenko, A., Edwards, A., Minor, W.(2012) FEBS J 279: 1093-1105
- PubMed: 22284390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08506.x
- Primary Citation Related Structures:
2QMO, 3MLE, 3QXC, 3QXH, 3QXJ, 3QXS, 3QXX, 3QY0 - PubMed Abstract:
Dethiobiotin synthetase (DTBS) is involved in the biosynthesis of biotin in bacteria, fungi, and plants. As humans lack this pathway, DTBS is a promising antimicrobial drug target. We determined structures of DTBS from Helicobacter pylori (hpDTBS) bound with cofactors and a substrate analog, and described its unique characteristics relative to other DTBS proteins. Comparison with bacterial DTBS orthologs revealed considerable structural differences in nucleotide recognition. The C-terminal region of DTBS proteins, which contains two nucleotide-recognition motifs, differs greatly among DTBS proteins from different species. The structure of hpDTBS revealed that this protein is unique and does not contain a C-terminal region containing one of the motifs. The single nucleotide-binding motif in hpDTBS is similar to its counterpart in GTPases; however, isothermal titration calorimetry binding studies showed that hpDTBS has a strong preference for ATP. The structural determinants of ATP specificity were assessed with X-ray crystallographic studies of hpDTBS·ATP and hpDTBS·GTP complexes. The unique mode of nucleotide recognition in hpDTBS makes this protein a good target for H. pylori-specific inhibitors of the biotin synthesis pathway.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, VA, USA.
Organizational Affiliation: