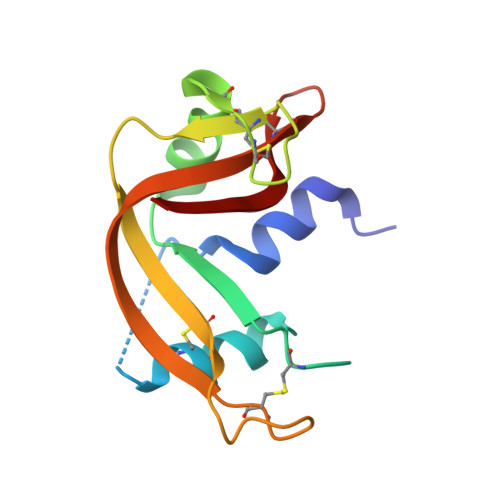
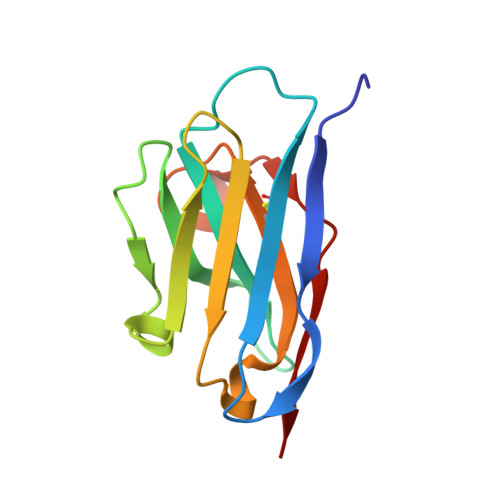
A combinatorial histidine scanning library approach to engineer highly pH-dependent protein switches.
Murtaugh, M.L., Fanning, S.W., Sharma, T.M., Terry, A.M., Horn, J.R.(2011) Protein Sci 20: 1619-1631
- PubMed: 21766385 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.696
- Primary Citation Related Structures:
3QSK - PubMed Abstract:
There is growing interest in the development of protein switches, which are proteins whose function, such as binding a target molecule, can be modulated through environmental triggers. Efforts to engineer highly pH sensitive protein-protein interactions typically rely on the rational introduction of ionizable groups in the protein interface. Such experiments are typically time intensive and often sacrifice the protein's affinity at the permissive pH. The underlying thermodynamics of proton-linkage dictate that the presence of multiple ionizable groups, which undergo a pK(a) change on protein binding, are necessary to result in highly pH-dependent binding. To test this hypothesis, a novel combinatorial histidine library was developed where every possible combination of histidine and wild-type residue is sampled throughout the interface of a model anti-RNase A single domain VHH antibody. Antibodies were coselected for high-affinity binding and pH-sensitivity using an in vitro, dual-function selection strategy. The resulting antibodies retained near wild-type affinity yet became highly sensitive to small decreases in pH, drastically decreasing their binding affinity, due to the incorporation of multiple histidine groups. Several trends were observed, such as histidine "hot-spots," which will help enhance the development of pH switch proteins as well as increase our understanding of the role of ionizable residues in protein interfaces. Overall, the combinatorial approach is rapid, general, and robust and should be capable of producing highly pH-sensitive protein affinity reagents for a number of different applications.
- Department of Chemistry and Biochemistry, Northern Illinois University, DeKalb, IL 60115, USA.
Organizational Affiliation: