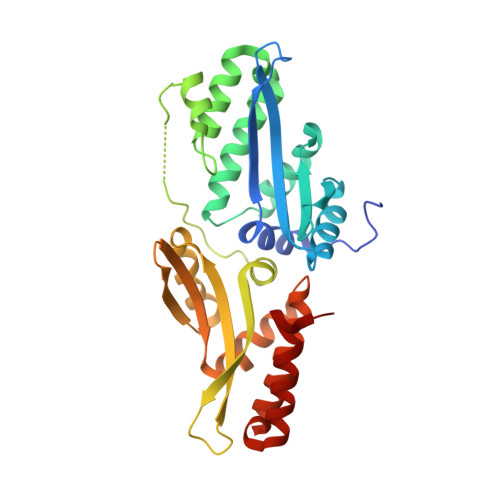
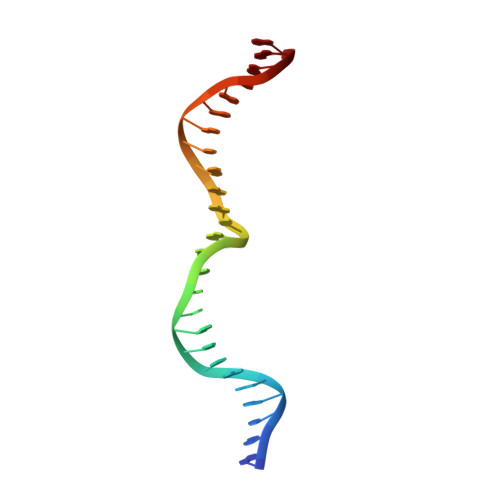
Tapping natural reservoirs of homing endonucleases for targeted gene modification.
Takeuchi, R., Lambert, A.R., Mak, A.N., Jacoby, K., Dickson, R.J., Gloor, G.B., Scharenberg, A.M., Edgell, D.R., Stoddard, B.L.(2011) Proc Natl Acad Sci U S A 108: 13077-13082
- PubMed: 21784983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1107719108
- Primary Citation Related Structures:
3QQY, 3R7P - PubMed Abstract:
Homing endonucleases mobilize their own genes by generating double-strand breaks at individual target sites within potential host DNA. Because of their high specificity, these proteins are used for "genome editing" in higher eukaryotes. However, alteration of homing endonuclease specificity is quite challenging. Here we describe the identification and phylogenetic analysis of over 200 naturally occurring LAGLIDADG homing endonucleases (LHEs). Biochemical and structural characterization of endonucleases from one clade within the phylogenetic tree demonstrates strong conservation of protein structure contrasted against highly diverged DNA target sites and indicates that a significant fraction of these proteins are sufficiently stable and active to serve as engineering scaffolds. This information was exploited to create a targeting enzyme to disrupt the endogenous monoamine oxidase B gene in human cells. The ubiquitous presence and diversity of LHEs described in this study may facilitate the creation of many tailored nucleases for genome editing.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue North A3-025, Seattle, WA 98109, USA.
Organizational Affiliation: