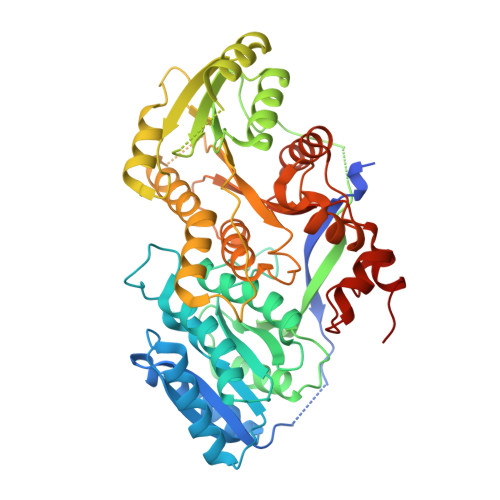
Structural and functional analysis of c2-type ketoreductases from modular polyketide synthases.
Zheng, J., Keatinge-Clay, A.T.(2011) J Mol Biology 410: 105-117
- PubMed: 21570406 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.04.065
- Primary Citation Related Structures:
3QP9 - PubMed Abstract:
The process by which α-stereocenters of polyketide intermediates are set by modular polyketide synthases (PKSs) when condensation is not immediately followed by reduction is mysterious. However, the reductase-incompetent ketoreductase (KR) from the third module of 6-deoxyerythronolide B synthase has been proposed to operate as a racemase, aiding in the epimerization process that reverses the orientation of the α-methyl group of the polyketide intermediate generated by the ketosynthase to the configuration observed in the 6-deoxyerythronolide B final product. To learn more about the epimerization process, the structure of the C2-type KR from the third module of the pikromycin synthase, analogous to the KR from the third module of 6-deoxyerythronolide B synthase, was determined to 1.88 Å resolution. This first structural analysis of this KR-type reveals differences from reductase-competent KRs such as that the site NADPH binds to reductase-competent KRs is occluded by side chains and the putative catalytic tyrosine possesses more degrees of freedom. The active-site geometry may enable C2-type KRs to align the thioester and β-keto groups of a polyketide intermediate to reduce the pK(a) of the α-proton and accelerate its abstraction. Results from in vivo assays of engineered PKSs support that C2-type KRs cooperate with epimer-specific ketosynthases to set the configurations of substituent-bearing α-carbons.
- Department of Chemistry and Biochemistry, University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: