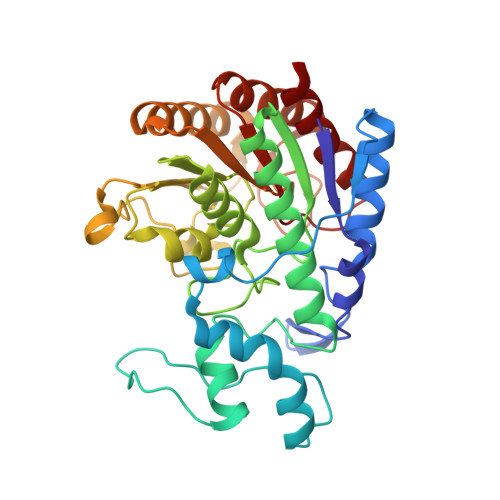
Structure of prokaryotic polyamine deacetylase reveals evolutionary functional relationships with eukaryotic histone deacetylases .
Lombardi, P.M., Angell, H.D., Whittington, D.A., Flynn, E.F., Rajashankar, K.R., Christianson, D.W.(2011) Biochemistry 50: 1808-1817
- PubMed: 21268586 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi101859k
- Primary Citation Related Structures:
3Q9B, 3Q9C, 3Q9E, 3Q9F - PubMed Abstract:
Polyamines are a ubiquitous class of polycationic small molecules that can influence gene expression by binding to nucleic acids. Reversible polyamine acetylation regulates nucleic acid binding and is required for normal cell cycle progression and proliferation. Here, we report the structures of Mycoplana ramosa acetylpolyamine amidohydrolase (APAH) complexed with a transition state analogue and a hydroxamate inhibitor and an inactive mutant complexed with two acetylpolyamine substrates. The structure of APAH is the first of a histone deacetylase-like oligomer and reveals that an 18-residue insert in the L2 loop promotes dimerization and the formation of an 18 Å long "L"-shaped active site tunnel at the dimer interface, accessible only to narrow and flexible substrates. The importance of dimerization for polyamine deacetylase function leads to the suggestion that a comparable dimeric or double-domain histone deacetylase could catalyze polyamine deacetylation reactions in eukaryotes.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104-6323, United States.
Organizational Affiliation: