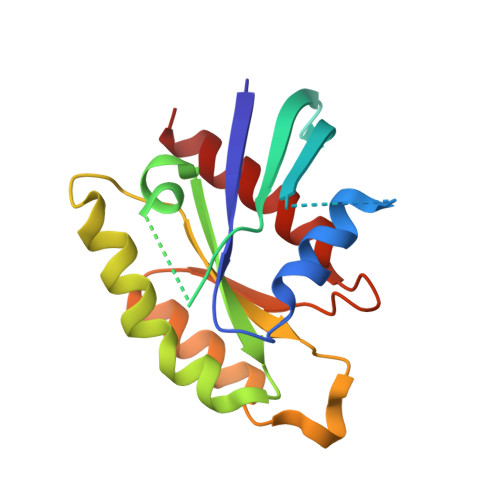
RGK Family G-Domain:GTP Analog Complex Structures and Nucleotide-Binding Properties.
Sasson, Y., Navon-Perry, L., Huppert, D., Hirsch, J.A.(2011) J Mol Biology 413: 372-389
- PubMed: 21903096 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.08.017
- Primary Citation Related Structures:
3Q72, 3Q7P, 3Q7Q, 3Q85 - PubMed Abstract:
The RGK family of small G-proteins, including Rad, Gem, Rem1, and Rem2, is inducibly expressed in various mammalian tissues and interacts with voltage-dependent calcium channels and Rho kinase. Many questions remain regarding their physiological roles and molecular mechanism. Previous crystallographic studies reported RGK G-domain:guanosine di-phosphate structures. To test whether RGK proteins undergo a nucleotide-induced conformational change, we determined the crystallographic structures of Rad:GppNHp and Rem2:GppNHp to 1.7 and 1.8 Å resolutions, respectively. Also, we characterized the nucleotide-binding properties and conformations for Gem, Rad, and several structure-based mutants using fluorescence spectroscopy. The results suggest that RGK G-proteins may not behave as Ras-like canonical nucleotide-induced molecular switches. Further, the RGK proteins have differing structures and nucleotide-binding properties, which may have implications for their varied action on effectors.
- Department of Biochemistry, Tel Aviv University, Ramat Aviv, Tel Aviv 69978, Israel.
Organizational Affiliation: