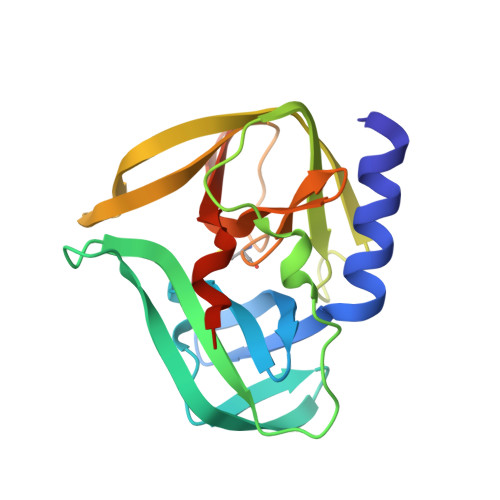
Structural Basis for Antiviral Inhibition of the Main Protease, 3C, from Human Enterovirus 93.
Costenaro, L., Kaczmarska, Z., Arnan, C., Janowski, R., Coutard, B., Sola, M., Gorbalenya, A.E., Norder, H., Canard, B., Coll, M.(2011) J Virol 85: 10764-10773
- PubMed: 21835784 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.05062-11
- Primary Citation Related Structures:
3Q3X, 3Q3Y, 3RUO - PubMed Abstract:
Members of the Enterovirus genus of the Picornaviridae family are abundant, with common human pathogens that belong to the rhinovirus (HRV) and enterovirus (EV) species, including diverse echo-, coxsackie- and polioviruses. They cause a wide spectrum of clinical manifestations ranging from asymptomatic to severe diseases with neurological and/or cardiac manifestations. Pandemic outbreaks of EVs may be accompanied by meningitis and/or paralysis and can be fatal. However, no effective prophylaxis or antiviral treatment against most EVs is available. The EV RNA genome directs the synthesis of a single polyprotein that is autocatalytically processed into mature proteins at Gln↓Gly cleavage sites by the 3C protease (3C(pro)), which has narrow, conserved substrate specificity. These cleavages are essential for virus replication, making 3C(pro) an excellent target for antivirus drug development. In this study, we report the first determination of the crystal structure of 3C(pro) from an enterovirus B, EV-93, a recently identified pathogen, alone and in complex with the anti-HRV molecules compound 1 (AG7404) and rupintrivir (AG7088) at resolutions of 1.9, 1.3, and 1.5 Å, respectively. The EV-93 3C(pro) adopts a chymotrypsin-like fold with a canonically configured oxyanion hole and a substrate binding pocket similar to that of rhino-, coxsackie- and poliovirus 3C proteases. We show that compound 1 and rupintrivir are both active against EV-93 in infected cells and inhibit the proteolytic activity of EV-93 3C(pro) in vitro. These results provide a framework for further structure-guided optimization of the tested compounds to produce antiviral drugs against a broad range of EV species.
- Institute for Research in Biomedicine, Barcelona, Spain.
Organizational Affiliation: