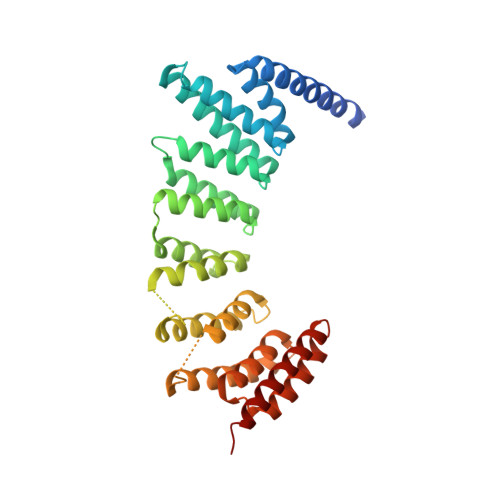
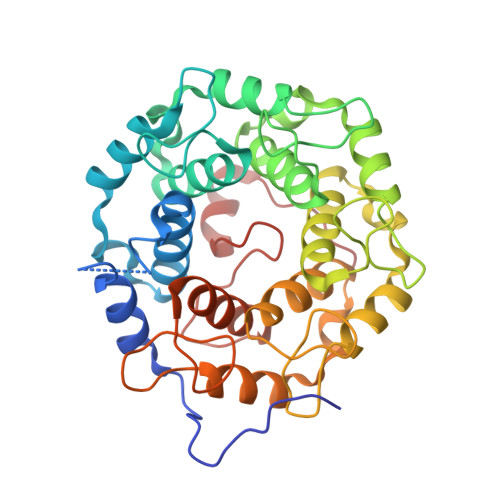
Structure-Guided Development of Selective RabGGTase Inhibitors.
Bon, R.S., Guo, Z., Stigter, E.A., Wetzel, S., Menninger, S., Wolf, A., Choidas, A., Alexandrov, K., Blankenfeldt, W., Goody, R.S., Waldmann, H.(2011) Angew Chem Int Ed Engl 50: 4957-4961
- PubMed: 21520375 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201101210
- Primary Citation Related Structures:
3PZ1, 3PZ2, 3PZ3, 3PZ4 - Max-Planck-Institut für molekulare Physiologie, Abt. Chemische Biologie, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: