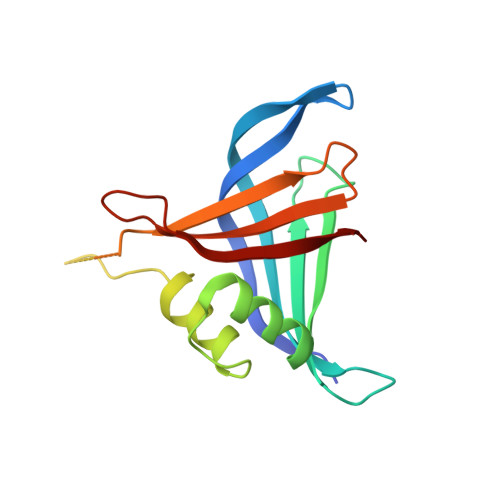
Structural basis of the 9-fold symmetry of centrioles.
Kitagawa, D., Vakonakis, I., Olieric, N., Hilbert, M., Keller, D., Olieric, V., Bortfeld, M., Erat, M.C., Fluckiger, I., Gonczy, P., Steinmetz, M.O.(2011) Cell 144: 364-375
- PubMed: 21277013 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2011.01.008
- Primary Citation Related Structures:
3PYI, 3Q0X, 3Q0Y - PubMed Abstract:
The centriole, and the related basal body, is an ancient organelle characterized by a universal 9-fold radial symmetry and is critical for generating cilia, flagella, and centrosomes. The mechanisms directing centriole formation are incompletely understood and represent a fundamental open question in biology. Here, we demonstrate that the centriolar protein SAS-6 forms rod-shaped homodimers that interact through their N-terminal domains to form oligomers. We establish that such oligomerization is essential for centriole formation in C. elegans and human cells. We further generate a structural model of the related protein Bld12p from C. reinhardtii, in which nine homodimers assemble into a ring from which nine coiled-coil rods radiate outward. Moreover, we demonstrate that recombinant Bld12p self-assembles into structures akin to the central hub of the cartwheel, which serves as a scaffold for centriole formation. Overall, our findings establish a structural basis for the universal 9-fold symmetry of centrioles.
- Swiss Institute for Experimental Cancer Research, School of Life Sciences, Swiss Federal Institute of Technology, EPFL, Lausanne, Switzerland.
Organizational Affiliation: