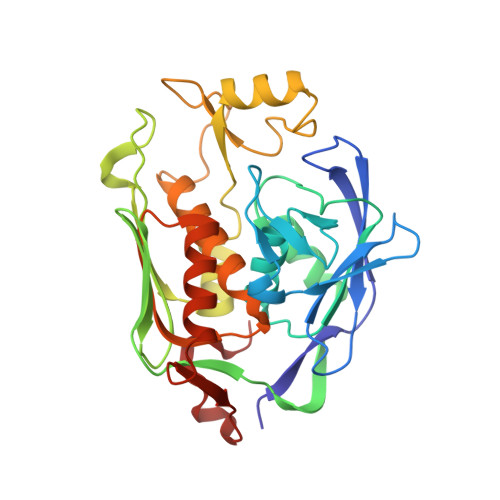
Syntheses, structures and antibiotic activities of LpxC inhibitors based on the diacetylene scaffold.
Liang, X., Lee, C.J., Chen, X., Chung, H.S., Zeng, D., Raetz, C.R., Li, Y., Zhou, P., Toone, E.J.(2011) Bioorg Med Chem 19: 852-860
- PubMed: 21194954 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmc.2010.12.017
- Primary Citation Related Structures:
3PS1, 3PS2, 3PS3 - PubMed Abstract:
Compounds inhibiting LpxC in the lipid A biosynthetic pathway are promising leads for novel antibiotics against multidrug-resistant Gram-negative pathogens. We report the syntheses and structural and biochemical characterizations of LpxC inhibitors based on a diphenyl-diacetylene (1,4-diphenyl-1,3-butadiyne) threonyl-hydroxamate scaffold. These studies provide a molecular interpretation for the differential antibiotic activities of compounds with a substituted distal phenyl ring as well as the absolute stereochemical requirement at the C2, but not C3, position of the threonyl group.
- Department of Chemistry, Jilin University, Changchun, PR China.
Organizational Affiliation: