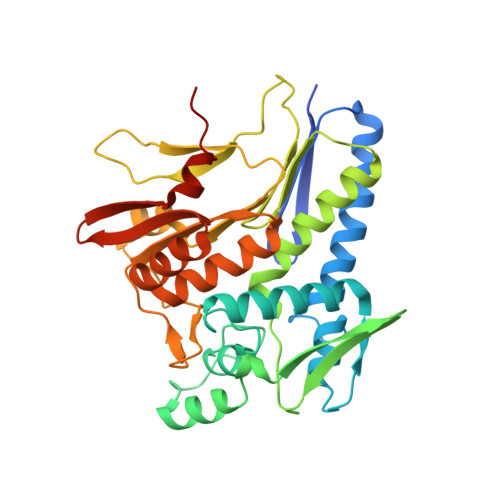
FAD binding by ApbE protein from Salmonella enterica: a new class of FAD-binding proteins.
Boyd, J.M., Endrizzi, J.A., Hamilton, T.L., Christopherson, M.R., Mulder, D.W., Downs, D.M., Peters, J.W.(2011) J Bacteriol 193: 887-895
- PubMed: 21148731 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00730-10
- Primary Citation Related Structures:
3PND - PubMed Abstract:
The periplasmic protein ApbE was identified through the analysis of several mutants defective in thiamine biosynthesis and was implicated as having a role in iron-sulfur cluster biosynthesis or repair. While mutations in apbE cause decreased activity of several iron-sulfur enzymes in vivo, the specific role of ApbE remains unknown. Members of the AbpE family include NosX and RnfF, which have been implicated in oxidation-reduction associated with nitrous oxide and nitrogen metabolism, respectively. In this work, we show that ApbE binds one FAD molecule per monomeric unit. The structure of ApbE in the presence of bound FAD reveals a new FAD-binding motif. Protein variants that are nonfunctional in vivo were generated by random and targeted mutagenesis. Each variant was substituted in the environment of the FAD and analyzed for FAD binding after reconstitution. The variant that altered a key tyrosine residue involved in FAD binding prevented reconstitution of the protein.
- Department of Bacteriology, University of Wisconsin, Madison, WI 53706, USA.
Organizational Affiliation: