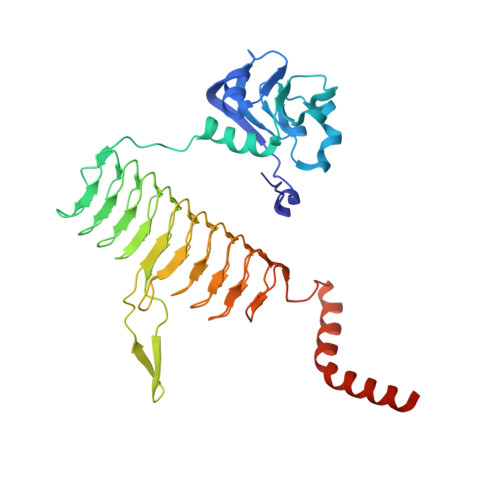
The structure of LpxD from Pseudomonas aeruginosa at 1.3 A resolution.
Badger, J., Chie-Leon, B., Logan, C., Sridhar, V., Sankaran, B., Zwart, P.H., Nienaber, V.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 749-752
- PubMed: 21795786 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111018811
- Primary Citation Related Structures:
3PMO - PubMed Abstract:
LpxD is a bacterial protein that is part of the biosynthesis pathway of lipid A and is responsible for transferring 3-hydroxymyristic acid from the R-3-hydroxymyristoyl-acyl carrier protein to the 2-OH group of UDP-3-O-(3-hydroxymyristoyl) glucosamine. The crystal structure of LpxD from Pseudomonas aeruginosa has been determined at high resolution (1.3 Å). The crystal belonged to space group H3, with unit-cell parameters a=b=106.19, c=93.38 Å, and contained one molecule in the asymmetric unit. The structure was solved by molecular replacement using the known structure of LpxD from Escherichia coli (PDB entry 3eh0) as a search model and was refined to Rwork=16.4% (Rfree=18.5%) using 91,655 reflections. The final protein model includes 355 amino-acid residues (including 16 amino acids from a 20 amino-acid N-terminal His tag), one chloride ion and two ethylene glycol molecules.
- Zenobia Therapeutics, 505 Coast Boulevard South, Suite 111, La Jolla, CA 92122, USA. john@zenobiatherapeutics.com
Organizational Affiliation: