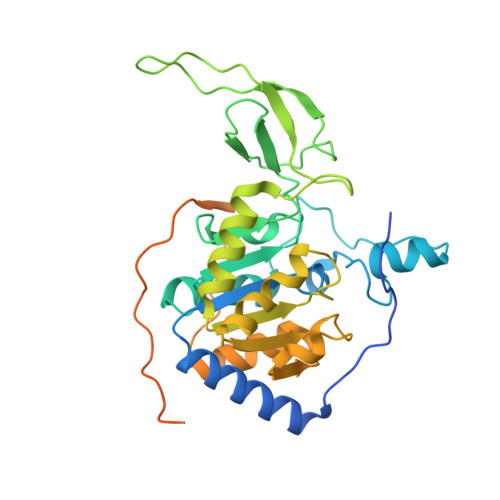
Structure and biochemical functions of SIRT6.
Pan, P.W., Feldman, J.L., Devries, M.K., Dong, A., Edwards, A.M., Denu, J.M.(2011) J Biological Chem 286: 14575-14587
- PubMed: 21362626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.218990
- Primary Citation Related Structures:
3K35, 3PKI, 3PKJ - PubMed Abstract:
SIRT6 is a member of the evolutionarily conserved sirtuin family of NAD(+)-dependent protein deacetylases and functions in genomic stability and transcriptional control of glucose metabolism. Early reports suggested that SIRT6 performs ADP-ribosylation, whereas more recent studies have suggested that SIRT6 functions mainly as a histone deacetylase. Thus, the molecular functions of SIRT6 remain uncertain. Here, we perform biochemical, kinetic, and structural studies to provide new mechanistic insight into the functions of SIRT6. Utilizing three different assays, we provide biochemical and kinetic evidence that SIRT6-dependent histone deacetylation produces O-acetyl-ADP-ribose but at a rate ∼1,000 times slower than other highly active sirtuins. To understand the molecular basis for such low deacetylase activity, we solved the first crystal structures of this class IV sirtuin in complex with ADP-ribose and the non-hydrolyzable analog of O-acetyl-ADP-ribose, 2'-N-acetyl-ADP-ribose. The structures revealed unique features of human SIRT6, including a splayed zinc-binding domain and the absence of a helix bundle that in other sirtuin structures connects the zinc-binding motif and Rossmann fold domain. SIRT6 also lacks the conserved, highly flexible, NAD(+)-binding loop and instead contains a stable single helix. These differences led us to hypothesize that SIRT6, unlike all other studied sirtuins, would be able to bind NAD(+) in the absence of an acetylated substrate. Indeed, we found that SIRT6 binds NAD(+) with relatively high affinity (K(d) = 27 ± 1 μM) in the absence of an acetylated substrate. Isothermal titration calorimetry and tryptophan fluorescence binding assays suggested that ADP-ribose and NAD(+) induce different structural perturbations and that NADH does not bind to SIRT6. Collectively, these new insights imply a unique activating mechanism and/or the possibility that SIRT6 could act as an NAD(+) metabolite sensor.
- Structural Genomics Consortium, University of Toronto, Toronto, Ontario M5G 1L7, Canada.
Organizational Affiliation: