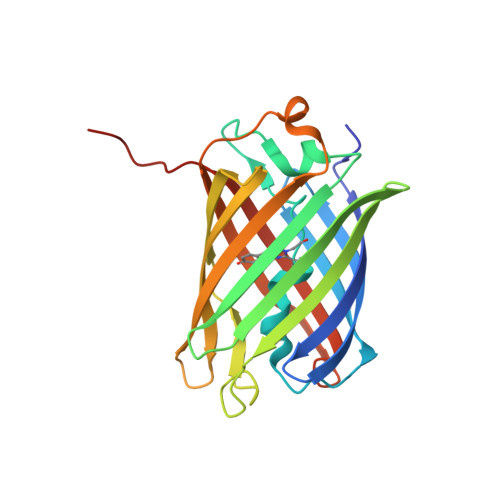
Crystallographic study of red fluorescent protein eqFP578 and its far-red variant Katushka reveals opposite pH-induced isomerization of chromophore.
Pletneva, N.V., Pletnev, V.Z., Shemiakina, I.I., Chudakov, D.M., Artemyev, I., Wlodawer, A., Dauter, Z., Pletnev, S.(2011) Protein Sci 20: 1265-1274
- PubMed: 21563226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.654
- Primary Citation Related Structures:
3PIB, 3PJ5, 3PJ7, 3PJB - PubMed Abstract:
The wild type red fluorescent protein eqFP578 (from sea anemone Entacmaea quadricolor, λ(ex) = 552 nm, λ(em) = 578 nm) and its bright far-red fluorescent variant Katushka (λ(ex) = 588 nm, λ(em) = 635 nm) are characterized by the pronounced pH dependence of their fluorescence. The crystal structures of eqFP578f (eqFP578 with two point mutations improving the protein folding) and Katushka have been determined at the resolution ranging from 1.15 to 1.85 Å at two pH values, corresponding to low and high level of fluorescence. The observed extinguishing of fluorescence upon reducing pH in eqFP578f and Katushka has been shown to be accompanied by the opposite trans-cis and cis-trans chromophore isomerization, respectively. Asn143, Ser158, His197 and Ser143, Leu174, and Arg197 have been shown to stabilize the respective trans and cis fluorescent states of the chromophores in eqFP578f and Katushka at higher pH. The cis state has been suggested as being primarily responsible for the observed far-red shift of the emission maximum of Katushka relative to that of eqFP578f.
- Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Science, Moscow 117997, Miklukho-Maklaya 16/10, Russia. pletnevs@mail.nih.gov
Organizational Affiliation: