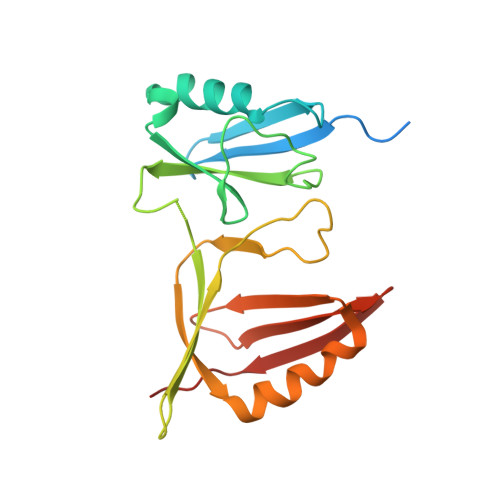
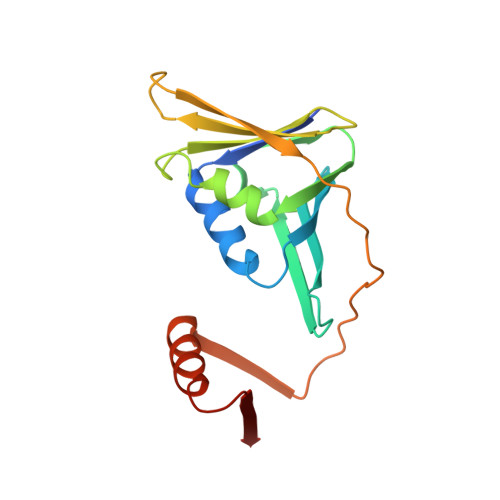
Crystal Structure of SUMO-Modified Proliferating Cell Nuclear Antigen.
Freudenthal, B.D., Brogie, J.E., Gakhar, L., Kondratick, C.M., Washington, M.T.(2011) J Mol Biology 406: 9-17
- PubMed: 21167178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.12.015
- Primary Citation Related Structures:
3PGE - PubMed Abstract:
Eukaryotic proliferating cell nuclear antigen (PCNA) is a replication accessory protein that functions in DNA replication, repair, and recombination. The various functions of PCNA are regulated by posttranslational modifications including mono-ubiquitylation, which promotes translesion synthesis, and sumoylation, which inhibits recombination. To understand how SUMO modification regulates PCNA, we generated a split SUMO-modified PCNA protein and showed that it supports cell viability and stimulates DNA polymerase δ activity. We then determined its X-ray crystal structure and found that SUMO occupies a position on the back face of the PCNA ring, which is distinct from the position occupied by ubiquitin in the structure of ubiquitin-modified PCNA. We propose that the back of PCNA has evolved to be a site of regulation that can be easily modified without disrupting ongoing reactions on the front of PCNA, such as normal DNA replication. Moreover, these modifications likely allow PCNA to function as a tool belt, whereby proteins can be recruited to the replication machinery via the back of PCNA and be held in reserve until needed.
- Department of Biochemistry, University of Iowa College of Medicine, Iowa City, IA 52242-1109, USA.
Organizational Affiliation: